Transfer RNA
Transfer RNA (tRNA) is a class of short-chain, non-coding ribonucleic acid (RNA) molecules in which each variety attaches to and transfers a specific amino acid to a polypeptide chain at the ribosomal site of protein synthesis during translation. As non-coding RNA, tRNA itself is not translated into a protein, but it does serve to transport amino acids for incorporation into proteins, using the chemical blueprint for the protein product provided by messenger RNA (mRNA), as derived from the DNA genes. Transfer RNA often is small, only about 74-79 nucleotides.
Each tRNA transports only one particular amino acid. This tRNA binds on one end to a specific codon (three-base region) in the mRNA and binds on the other end to the amino acid specified by that codon. As a result, it places that amino acid in the correct sequence in the growing polypeptide according to the template (sequence of nucleotides) provided by the mRNA (Alberts et al. 1989).
The intricacy and complex coordination of the process of building a protein utilizing tRNA is striking. The mechanism is dependent on each tRNA binding with each amino acid to have a precise three-dimensional structure, requires a pairing up of the tRNA anticodon with the mRNA codon, and is dependent on specific enzymes for linking each amino acid with the necessary tRNA, among other precise structures and processes.
Overview
A protein is a complex, high-molecular mass organic compound comprising amino acids joined together in chains. The site of protein synthesis is the ribosome. It is here that messenger RNA's (mRNA) provide the code for linking amino acids together to form new proteins and where transfer RNAs (tRNA) transfer specific amino acids to the growing polypeptide chain during the translation of the mRNA into a protein. The chemical blueprint for the protein product is provided by the mRNA.
Transfer RNA has a 3' terminal site for the amino acid attachment. The amino acid covalently links by its carboxyl end to this 3' end. This covalent linkage is catalyzed by enzymes called aminoacyl tRNA synthetases, with a different enzyme for each amino acid (Alberts et al. 1989). The particular synthetase enzyme couples the amino acid to the appropriate tRNA molecule.
The tRNA also contains a three base region called the anticodon that can base pair to the corresponding three base codon region on mRNA. Each type of tRNA molecule can be attached to only one type of amino acid; however, because the genetic code contains multiple codons that specify the same amino acid, tRNA molecules bearing different anticodons may carry the same amino acid. Most amino acids have several tRNAs with which they can bind.
The linkage of the tRNA to a specific codon of mRNA and the linkage of the tRNA to a specific amino acid results in the amino acid being placed in the appropriate place in the growing polypeptide chain.
The linkage of the tRNA-amino acid at the carboxyl end of the amino acid is a high-energy linkage and thus the amino acid is "activated" so that it can react readily with the amino group of the next amino acid to be added in the sequence, forming a peptide bond.
Structure
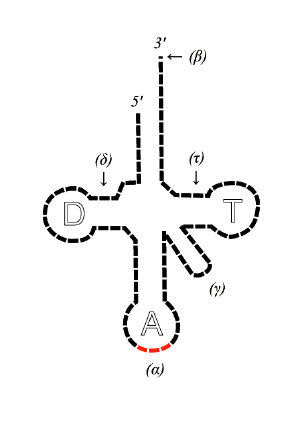
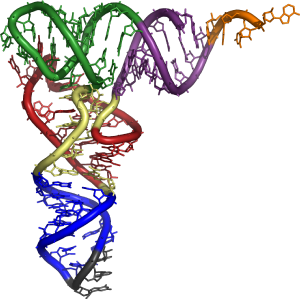
Transfer RNA has primary structure, secondary structure (usually visualized as the cloverleaf structure), and tertiary structure. (All tRNAs have a similar L-shaped 3D structure that allows them to fit into the P and A sites of the ribosome.)
- The 5'-terminal phosphate group.
- The acceptor stem is a 7-bp stem made by the base pairing of the 5'-terminal nucleotide with the 3'-terminal nucleotide (which contains the CCA 3'-terminal group used to attach the amino acid). The acceptor stem may contain non-Watson-Crick base pairs.
- The CCA tail is a CCA sequence at the 3' end of the tRNA molecule. This sequence is important for the recognition of tRNA by enzymes critical in translation. In prokaryotes, the CCA sequence is transcribed. In eukaryotes, the CCA sequence is added during processing and therefore does not appear in the tRNA gene.
- The D arm is a 4 bp stem ending in a loop that often contains dihydrouridine.
- The anticodon arm is a 5-bp stem whose loop contains the anticodon.
- The T arm is a 5 bp stem containing the sequence TΨC where Ψ is a pseudouridine.
- Bases that have been modified, especially by methylation, occur in several positions outside the anticodon. The first anticodon base is sometimes modified to inosine (derived from adenine) or pseudouridine (derived from uracil).
Anticodon
An anticodon (Felsenfeld and Cantoni 1964) is a unit made up of three nucleotides that correspond to the three bases of the codon on the mRNA. Each tRNA contains a specific anticodon triplet sequence that can base-pair to one or more codons for an amino acid. For example, one codon for lysine is AAA; the anticodon of a lysine tRNA might be UUU. Some anticodons can pair with more than one codon due to a phenomenon known as wobble base pairing. Frequently, the first nucleotide of the anticodon is one of two not found on mRNA: inosine and pseudouridine, which can hydrogen bond to more than one base in the corresponding codon position. In the genetic code, it is common for a single amino acid to be specified by all four third-position possibilities; for example, the amino acid glycine is coded for by the codon sequences GGU, GGC, GGA, and GGG.
To provide a one-to-one correspondence between tRNA molecules and codons that specify amino acids, 61 tRNA molecules would be required per cell. However, many cells contain fewer than 61 types of tRNAs because the wobble base is capable of binding to several, though not necessarily all, of the codons that specify a particular amino acid (Lodish et al. 2007).
Aminoacylation
Aminoacylation is the process of adding an aminoacyl group to a compound. It produces tRNA molecules with their CCA 3' ends covalently linked to an amino acid.
Each tRNA is aminoacylated (or charged) with a specific amino acid by an aminoacyl tRNA synthetase. There is normally a single aminoacyl tRNA synthetase for each amino acid, despite the fact that there can be more than one tRNA, and more than one anticodon, for an amino acid. Recognition of the appropriate tRNA by the synthetases is not mediated solely by the anticodon, and the acceptor stem often plays a prominent role.
Reaction:
- amino acid + ATP → aminoacyl-AMP + PPi
- aminoacyl-AMP + tRNA → aminoacyl-tRNA + AMP
Binding to ribosome
The ribosome has three binding sites for tRNA molecules: The A, P, and E sites. During translation, the A site binds an incoming aminoacyl-tRNA as directed by the codon currently occupying this site. This codon specifies the next amino acid to be added to the growing peptide chain. The A site only works after the first aminoacyl-tRNA has attached to the P site. The P-site codon is occupied by peptdyl-tRNA that is a tRNA with multiple amino acids attached as a long chain. The P site is actually the first to bind to aminoacyl tRNA. This tRNA in the P site carries the chain of amino acids that has already been synthesized. The E site is occupied by the empty tRNA as it is about to exit the ribosome.
Transfer RNA genes
Organisms vary in the number of tRNA genes in their genome. The nematode worm C. elegans, a commonly used model organism in genetics studies, has 29,647 genes in its nuclear genome (WormBase 2008), of which 620 code for tRNA (Spieth and Lawson 2006; Hartwell et al. 2004). The budding yeast Saccharomyces cerevisiae has 275 tRNA genes in its genome. In the human genome, which according to current estimates has about 27,161 genes in total (Ensembl 2008), there are about 4,421 non-coding RNA genes, which include tRNA genes. There are 22 mitochondrial tRNA genes (Ensembl 2008); 497 nuclear genes encoding cytoplasmic tRNA molecules, and there are 324 tRNA-derived putative pseudogenes (Lander et al. 2001).
Cytoplasmic tRNA genes can be grouped into 49 families according to their anticodon features. These genes are found on all chromosomes, except 22 and Y chromosome. High clustering on 6p is observed (140 tRNA genes), as well on 1 chromosome (Lander et al. 2001).
Transfer RNA molecules are transcribed (in eukaryotic cells) by RNA polymerase III (White 1997), unlike messenger RNA, which is transcribed by RNA polymerase II. Pre-tRNAs contain introns; in bacteria these self-splice, whereas in eukaryotes and archaea they are removed by tRNA splicing endonuclease (Abelson et al. 1998).
History
The existence of tRNA was first hypothesized by Francis Crick, based on the assumption that there must exist an adapter molecule capable of mediating the translation of the RNA alphabet into the protein alphabet. Significant research on structure was conducted in the early 1960s, by Alex Rich and Don Caspar, two researchers in Boston, the Jacques Fresco group in Princeton University, and a United Kingdom group at King's College London (Clark 2006). A later publication reported the primary structure in 1965, by Robert W. Holley. The secondary and tertiary structures were derived from X-ray crystallography studies reported independently in 1974, by American and British research groups headed, respectively, by Alexander Rich and Aaron Klug.
ReferencesISBN links support NWE through referral fees
- Abelson, J., C. R. Trotta, and H. Li. 1998. tRNA splicing. J Biol Chem 273(21): 12685–12688. PMID 9582290. Retrieved October 5, 2008.
- Alberts, B., D. Bray, J. Lewis, M. Raff, K. Roberts, and J. D. Watson. Molecular Biology of the Cell, 2nd edition. New York: Garland Publishing, 1989. ISBN 0824036956.
- Alberts, B., A. Johnson, J. Lewis, M. Raff, K. Roberts, and P. Walter. 2002. Molecular Biology of the Cell, 4th edition. New York: Garland Science. ISBN 0815332181.
- Clark, B. F. C. 2006. The crystal structure of tRNA. J. Biosci. 31(4): 453–7. PMID 17206065. Retrieved October 5, 2008.
- Ensembl. 2008. Explore the Homo sapiens genome. Ensemble Release 50. Retrieved October 5, 2008.
- Felsenfeld, G., and G. Cantoni. 1964. Use of thermal denaturation studies to investigate the base sequence of yeast serine sRNA. Proc Natl Acad Sci U S A 51: 818–26. PMID 14172997. Retrieved October 5, 2008.
- Hartwell, L. H., L. Hood, M. L. Goldberg, A. E. Reynolds, L. M. Silver, and R. C. Veres. 2004. Genetics: From Genes to Genomes, 2nd edited. New York: McGraw-Hill. ISBN 0072919302.
- Lander, E., L. M. Linton, B. Birren, et al. 2001. Initial sequencing and analysis of the human genome. Nature 409(6822): 860–921. PMID 11237011. Retrieved October 5, 2008.
- Lodish, H., A. Berk, C. A. Kaiser, M. Krieger, M. P. Scott, A. Bretscher, H. Ploegh, and P. Matsudaira. 2007. Molecular Biology of the Cell. New York: WH Freeman. ISBN 9780716776017.
- Spieth, J., and D. Lawson. 2006. Overview of gene structure (January 18, 2006). WormBook: The C. elegans Research Community. Retrieved October 5, 2008.
- White, R. J. 1997. Regulation of RNA polymerases I and III by the retinoblastoma protein: A mechanism for growth control? Trends in Biochemical Sciences 22(3): 77–80. Retrieved October 5, 2008.
- Wormbase. 2008. Wormbase release WS187. Wormbase. Retrieved October 5, 2008.
External links
All links retrieved May 1, 2023.
Credits
New World Encyclopedia writers and editors rewrote and completed the Wikipedia article in accordance with New World Encyclopedia standards. This article abides by terms of the Creative Commons CC-by-sa 3.0 License (CC-by-sa), which may be used and disseminated with proper attribution. Credit is due under the terms of this license that can reference both the New World Encyclopedia contributors and the selfless volunteer contributors of the Wikimedia Foundation. To cite this article click here for a list of acceptable citing formats.The history of earlier contributions by wikipedians is accessible to researchers here:
The history of this article since it was imported to New World Encyclopedia:
Note: Some restrictions may apply to use of individual images which are separately licensed.