Translation (biology)
In biology, translation is the cellular process in which proteins are produced by decoding, or translating, particular genetic information of the DNA using a messenger RNA (mRNA) intermediate as the template. Also known as protein synthesis or protein biosynthesis, translation occurs in the cytoplasm where the ribosomes are located and utilizes transfer RNAs (tRNAs) for attaching the specific amino acids that make up the protein. Translation is the second of two basic steps in the process of converting genes to proteins, with the first step being the transcription of a portion of the DNA into the mRNA. Then during translation, the mRNA guides the assembly of the amino acids into the particular sequence.
Translation proceeds in four phases: Activation, initiation, elongation, and termination, all describing the growth of the amino acid chain, or polypeptide, that is the product of translation. In reflecting the remarkable harmony of nature, this is a very complex and yet stunningly precise process, involving numerous components, movements, and reactions that are intricately coordinated.
The capacity of disabling or inhibiting translation in protein biosynthesis is used by antibiotics such as: anisomycin, cycloheximide, chloramphenicol, tetracycline, streptomycin, erythromycin, puromycin, and so forth. Prokaryotic ribosomes have a different structure than eukaryotic ribosomes, and thus antibiotics can specifically target bacterial infections without any detriment to the host's cells.
Basic process
This article is part of the series on: Gene expression | |||
| Introduction to Genetics | |||
| General flow: DNA > RNA > Protein | |||
| special transfers (RNA > RNA, RNA > DNA, Protein > Protein) | |||
| Genetic code | |||
| Transcription | |||
| Transcription (Transcription factors, RNA Polymerase,promoter) | |||
| post-transcriptional modification (hnRNA,Splicing) | |||
| Translation | |||
| Translation (Ribosome,tRNA) | |||
| post-translational modification (functional groups, peptides, structural changes) | |||
| gene regulation | |||
| epigenetic regulation (Hox genes, Genomic imprinting) | |||
| transcriptional regulation | |||
| post-transcriptional regulation (sequestration, alternative splicing,miRNA) | |||
| post-translational regulation (reversible,irrevesible) | |||
The conversion of genes to proteins is essentially a two-step process: Transcription and translation. Sometimes the term "protein synthesis" is used to refer only to protein translation, since this is the first stage in actually building the protein, but the entire process of expressing a gene into a protein requires transcription as well.
Proteins are composed of amino acids joined together in chains. The ultimate template for construction of a protein is the DNA of the organism. However, the site of protein synthesis is the ribosome and it is messenger RNA's (mRNA) that provide the code or chemical blueprint for linking amino acids together to form new proteins. Messenger RNAs are synthesized from the DNA template in the process known as DNA transcription and then carry this coding information to the ribosomes, where the translation into proteins takes place.
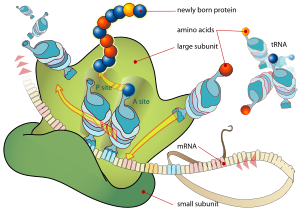
The ribosome is a multi-subunit structure containing rRNA and proteins. It is the "factory" where amino acids are assembled into proteins. Ribosomes are made of a small and large subunit that surrounds the mRNA.
The mRNA that codes for a protein is a direct copy of the sequence of nucleotides corresponding to a specific region (gene) in one of the two complementary DNA strands. In mRNA, as in DNA, genetic information is encoded in the sequence of four nucleotides arranged into codons of three bases each. Each codon encodes for a specific amino acid, except the stop codons that terminate protein synthesis. With four different nucleotides, there are 64 different codons possible. All but three of these combinations (UAA, UGA, and UAG) code for a particular amino acid. However, there are only twenty amino acids, so some amino acids are specified by more than one codon (Zengel 2003).
Transfer RNAs (tRNAs) transport amino acids to the ribosomes and then act to transfer the correct amino acid to the correct part of the growing polypeptide. Transfer RNAs are small noncoding RNA chains (74-93 nucleotides). They have a site for amino acid attachment, and a site called an anticodon. The anticodon is an RNA triplet complementary to the mRNA triplet that codes for their cargo amino acid. At the site of protein synthesis, tRNAs bind on one end to specific codons (three-base region) in the mRNA and bind on the other end to the amino acids specified by that codon. Transfer RNAs thus place the amino acids in the correct sequence in the growing polypeptide according to the template (sequence of nucleotides) provided by the mRNA, as derived from the DNA gene (Alberts et al. 1989). That is, the nucleic acid polymer is translated into a protein. Each tRNA transports only one particular amino acid.
Thus, for example, since adenine (A) pairs with uracil (U) and guanine (G) pairs with cytosine (C), a tRNA with a UUU anticodon recognizes the AAA codon of mRNA and brings its amino acid (lysine in this case) to add to the protein at that part of the mRNA (Zengel 2003).
Aminoacyl tRNA synthetase (an enzyme) catalyzes the bonding between specific tRNAs and the amino acids that their anticodons sequences call for. The product of this reaction is an aminoacyl-tRNA molecule. This aminoacyl-tRNA travels inside the ribosome, where mRNA codons are matched through complementary base pairing to specific tRNA anticodons. The amino acids that the tRNAs carry are then used to assemble a protein.
The energy required for translation of proteins is significant. For a protein containing n amino acids, the number of high-energy Phosphate bonds required to translate it is 4n-1.
Four phases of translation
Translation proceeds in four phases: Activation, initiation, elongation, and termination.
In activation, the correct amino acid is covalently bonded to the correct transfer RNA (tRNA). While this is not technically a step in translation, it is required for translation to proceed. The amino acid is joined by its carboxyl group to the 3' OH of the tRNA by an ester bond. When the tRNA has an amino acid linked to it, it is termed "charged."
The first true stage of translation is initiation. Initiation involves the small subunit of the ribosome binding to 5' (five prime) end of mRNA with the help of initiation factors (IF). RNA molecules, like DNA molecules, have directionality, which is indicated by reference to either the 5’ end or the 3’ (three prime) end. The code is read in the 5’ to 3’ direction (Zengel 2003).
During elongation, additional amino acids are added progressively to the chain. The new amino acid is attached as a new tRNA brings the next amino acid and is based paired by its anticodon ot the next codon in the mRNA and then the new amino acid is paired to the previous amino acid by a peptide bond. This step is carried out by the ribosome. As amino acids are added, the mRNA slides along the ribosome, three bases at a time. The earlier tRNA (and initially the special initiator RNA) sits on the site of the ribosome called the peptidyl or P site and the new tRNA uses the acceptor or A site (Zengel 2003).
Termination of the polypeptide happens when the A site of the ribosome faces a stop codon (UAA, UAG, or UGA). When this happens, no tRNA can recognize it, but a releasing factor can recognize nonsense codons and causes the release of the polypeptide chain.
The 5' end of the mRNA gives rise to the proteins N-terminal and the direction of translation can therefore be stated as N->C.
Prokaryotic versus eukaryotic translation
Prokaryotic translation and eukaryotic translation exhibit differences in various aspects of the process. One of these differences relates to the AUG codon. In the initiation step of translation, a special initiator tRNA binds to a special site on the small subunit of the ribosome. The mRNA also is loaded on and positioned so that the initiation codon, usually AUG, is based paired with the anticodon of this tRNA. AUG codes for the amino acid methionine, but some AUGs code for methionines within the polypeptide chain, not just at the initiation site. The difference in these AUG codon’s must be recognized. In bacteria, there is additional information contained within the mRNA sequence right before the initiating AUG, known as the Shine-Dalgarno sequence, that helps in this recognition. In eukaryote cells, there is a different strategy, where the mRNA contains a special structure at its 5’ end (Zengel 2003).
In eukaryotes, mRNA precursors must be processed in the nucleus (such as capping, polyadenylation, splicing) before they are exported to the cytoplasm for translation.
The process of termination is similar in eukaryotes and prokaryotes.
The rate of translation varies; it is significantly higher in prokaryotic cells (up to 17 to 21 amino acid residues per second) than in eukaryotic cells (up to 6 to 7 amino acid residues per second) (Ross and Orlowski 1982).
Translation by hand and by computer
It is possible to translate either by hand (for short sequences) or by computer (after first programming one appropriately); this allows biologists and chemists to draw out the chemical structure of the encoded protein on paper.
Translation by hand
In translation by hand, one first converts each template DNA base to its RNA complement (note that the complement of A is now U), as shown below. Note that the template strand of the DNA is the one the RNA is polymerized against; the other DNA strand would be the same as the RNA, but with thymine instead of uracil, since uracil is used in place of thymine in RNA.
DNA -> RNA A -> U T -> A G -> C C -> G
One then splits the RNA into triplets (groups of three bases). Note that there are 3 translation "windows," or reading frames, depending on where you start reading the code.
Finally, one uses a genetic code table to translate the above into a structural formula as used in chemistry.
This process yields the primary structure of the protein. However, proteins tend to fold, depending in part on hydrophilic and hydrophobic segments along the chain. Secondary structure can often still be guessed at, but the proper tertiary structure is often very hard to determine.
This approach may not give the correct amino acid composition of the protein, in particular if unconventional amino acids such as selenocysteine are incorporated into the protein, which is coded for by a conventional stop codon in combination with a downstream hairpin (SElenoCysteine Insertion Sequence, or SECIS).
Translation by computer
Many computer programs capable of translating a DNA/RNA sequence into protein sequence exist. Normally, this is performed using the Standard Genetic Code; many bioinformaticians have written at least one such program at some point in their education. However, few programs can handle all the "special" cases, such as the use of the alternative initiation codons. For example, the rare alternative start codon CTG codes for methionine when used as a start codon, and for leucine in all other positions.
Example: Condensed translation table for the Standard Genetic Code (from the NCBI Taxonomy webpage).
AAs = FFLLSSSSYY**CC*WLLLLPPPPHHQQRRRRIIIMTTTTNNKKSSRRVVVVAAAADDEEGGGG Starts = ---M---------------M---------------M---------------------------- Base1 = TTTTTTTTTTTTTTTTCCCCCCCCCCCCCCCCAAAAAAAAAAAAAAAAGGGGGGGGGGGGGGGG Base2 = TTTTCCCCAAAAGGGGTTTTCCCCAAAAGGGGTTTTCCCCAAAAGGGGTTTTCCCCAAAAGGGG Base3 = TCAGTCAGTCAGTCAGTCAGTCAGTCAGTCAGTCAGTCAGTCAGTCAGTCAGTCAGTCAGTCAG
Translation tables
Even when working with ordinary eukaryotic sequences such as the yeast genome, it is often desired to be able to use alternative translation tables— namely for translation of the mitochondrial genes. Currently the following translation tables are defined by the NCBI Taxonomy Group for the translation of the sequences in GenBank:
1: The Standard 2: The Vertebrate Mitochondrial Code 3: The Yeast Mitochondrial Code 4: The Mold, Protozoan, and Coelenterate Mitochondrial Code and the Mycoplasma/Spiroplasma Code 5: The Invertebrate Mitochondrial Code 6: The Ciliate, Dasycladacean and Hexamita Nuclear Code 9: The Echinoderm and Flatworm Mitochondrial Code 10: The Euplotid Nuclear Code 11: The Bacterial and Plant Plastid Code 12: The Alternative Yeast Nuclear Code 13: The Ascidian Mitochondrial Code 14: The Alternative Flatworm Mitochondrial Code 15: Blepharisma Nuclear Code 16: Chlorophycean Mitochondrial Code 21: Trematode Mitochondrial Code 22: Scenedesmus obliquus mitochondrial Code 23: Thraustochytrium Mitochondrial Code
ReferencesISBN links support NWE through referral fees
- Alberts, B., D. Bray, J. Lewis, M. Raff, K. Roberts, and J. D. Watson. 1989. Molecular Biology of the Cell, 2nd edition. New York: Garland Publishing. ISBN 0824036956.
- Champe, P. C., R. A Harvey, and D. R. Ferrier. 2005. Lippincott's Illustrated Reviews: Biochemistry, 3rd edition. Lippincott Williams & Wilkins. ISBN 0781722659.
- Nelson, D. L. and M. M. Cox. 2005. Lehninger Principles of Biochemistry, 4th edition. W. H. Freeman. ISBN 0716743396.
- Ross, J. F., and M. Orlowski. 1982. Growth-rate-dependent adjustment of ribosome function in chemostat-grown cells of the fungus Mucor racemosus. J Bacteriol. 149(2): 650–653. PMCID: PMC216554.
- Zengel, J. 2003. Translation. In R. Robinson, Genetics. New York: Macmillan Reference USA. OCLC 55983868.
| |||||||||||
| ||||||||
Credits
New World Encyclopedia writers and editors rewrote and completed the Wikipedia article in accordance with New World Encyclopedia standards. This article abides by terms of the Creative Commons CC-by-sa 3.0 License (CC-by-sa), which may be used and disseminated with proper attribution. Credit is due under the terms of this license that can reference both the New World Encyclopedia contributors and the selfless volunteer contributors of the Wikimedia Foundation. To cite this article click here for a list of acceptable citing formats.The history of earlier contributions by wikipedians is accessible to researchers here:
The history of this article since it was imported to New World Encyclopedia:
Note: Some restrictions may apply to use of individual images which are separately licensed.