Recombinant DNA
Recombinant DNA is a form of genetically engineered DNA that is created by taking DNA strands from one organism and combining or inserting these strands into the DNA of a host organism of a different species. What is yielded is a form of artificial DNA, combining DNA sequences that would not normally occur together (Berg et al. 2007). It differs from genetic recombination, in that it does not occur through processes within the cell or ribosome, but is exclusively engineered (Berg et al. 2007).
Recombinant DNA technology allows scientists to transplant a gene for a particular protein into a rapidly reproducing bacteria to mass produce the protein or insert it into the genome of a crop to provide a certain benefit. As a result of this technology, bacteria have been used to produce human insulin beginning in 1978. Additional therapies and production of drugs have utilized recombinant DNA techniques since then, and the technology is gaining more and more application in pharmacology and agriculture. Along with these advances are concerns about the safety of recombinant DNA technologies and the potential for its misuse.
Overview
DNA, or deoxyribonucleic acid, is a nucleic acid molecule that contains the genetic instructions used in the development and functioning of all living organisms. The main role of DNA is the long-term storage of information and it is often compared to a set of blueprints, since DNA contains the instructions needed to construct other components of cells, such as proteins and RNA molecules. The DNA segments that carry this genetic information are called genes, but other DNA sequences have structural purposes, or are involved in regulating the use of this genetic information.
Chemically, DNA is a long polymer of simple units called nucleotides, with a backbone made of sugars and phosphate atoms joined by ester bonds. Attached to each sugar is one of four types of molecules, called nucleobases. It is the sequence of these four bases along the backbone that encodes information. This information is read using the genetic code, which specifies the sequence of the amino acids within proteins. The code is read by copying stretches of DNA into the related nucleic acid RNA, in a process called transcription. Most of these RNA molecules are used to synthesize proteins, but others are used directly in structures such as ribosomes and spliceosomes.
Recombinant DNA is a form of artificial DNA which is engineered through the combination or insertion of one or more DNA strands (Berg et al. 2007). In terms of genetic modification, recombinant DNA is produced through the addition of relevant DNA into an existing organismal genome, such as the plasmid of bacteria, to code for or alter different traits for a specific purpose, such as immunity (Berg et al. 2007).
The Recombinant DNA technique was engineered by Stanley Norman Cohen and Herbert Boyer in 1973. They published their findings in a 1974 paper entitled, "Construction of Biologically Functional Bacterial Plasmids in vitro," which described a technique to isolate and amplify genes or DNA segments and insert them into another cell with precision, creating a transgenic bacterium. Recombinant DNA technology was made possible by the discovery of restriction endonucleases (an enzyme that cuts double-stranded DNA) by Werner Arber, Daniel Nathans, and Hamilton Smith, for which they received the 1978 Nobel Prize in Medicine.
Because of the importance of DNA in the replication of new structures and characteristics of living organisms, it has widespread importance in recapitulating via viral or non-viral vectors, both desirable and undesirable characteristics of a species to achieve characteristic change or to counteract effects caused by genetic or imposed disorders that have effects upon cellular or organismal processes (Kaplan et al. 1980). Through the use of recombinant DNA, genes that are identified as important can be amplified and isolated for use in other species or applications, where there may be some form of genetic illness or discrepancy, and provides a different approach to complex biological problem solving (Kaplan et al. 1980).
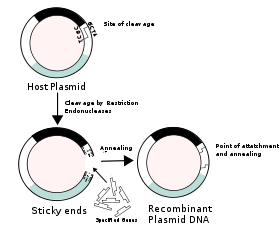
The basic technique for creating recombinant DNA involves using a restriction enzyme to cut DNA at specific sites in a vector DNA. A restriction enzyme (or restriction endonuclease) is an enzyme that cuts double-stranded DNA, like a molecular scissors. The enzyme makes two incisions, one through each of the sugar-phosphate backbones (i.e., each strand) of the double helix without damaging the nitrogenous bases. DNA of a host organism is similarly treated with the restriction enzyme, and then the two DNA segments are joined, suing another enzyme, to make an intact DNA molecule.
Applications and methods
Cloning and relation to plasmids
The use of cloning is interrelated with Recombinant DNA in classical biology, as the term "clone" refers to a cell or organism derived from a parental organism, with modern biology referring to the term as a collection of cells derived from the same cell that remain identical (Berg et al. 2007). In the classical instance, the use of recombinant DNA provides the initial cell from which the host organism is then expected to recapitulate when it undergoes further cell division—with bacteria remaining a prime example due to the use of viral vectors in medicine, which contain recombinant DNA inserted into a structure known as a plasmid (Berg et al. 2007).
Plasmids are extrachromosomal, self replicating, circular forms of DNA present in most bacteria, such as Escherichia coli (E. Coli), contain genes related to catabolism and metabolic activity, and allow the carrier bacterium to survive and reproduce in conditions present within other species and environments (Berg et al. 2007). These genes represent characteristics of resistance to bacteriophages and antibiotics and some heavy metals, but can also be fairly easily removed or separated from the plasmid by restriction endonucleases (Berg et al. 2007). Restriction enzymes regularly produce "sticky ends" and allow the attachment of a selected segment of DNA, which codes for more "reparative" substances, such as peptide hormone medications, including insulin, growth hormone, and oxytocin (Berg et al. 2007). In the introduction of useful genes into the plasmid, the bacteria is then used as a viral vector, which is encouraged to reproduce so as to recapitulate the altered DNA within other cells it infects and increase the amount of cells with the recombinant DNA present within them.
The use of plasmids is also key within gene therapy, where their related viruses are used as cloning vectors or carriers, which are means of transporting and passing on genes in recombinant DNA through viral reproduction throughout an organism (Berg et al. 2007). As a general definition of plasmids, the definition is that they contain three common features: A replicator, a selectable marker, and a cloning site (Berg et al. 2007). The replicator, or "ori," refers to the origin of replication with regards to location and bacteria where replication begins (Berg et al. 2007). The marker refers to a gene that usually contains resistance to an antibiotic, but may also refer to a gene that is attached alongside the desired one, such as that which confers luminescence to allow identification of successfully recombined DNA (Berg et al. 2007). The cloning site is a sequence of nucleotides representing one or more positions where cleavage by restriction endonucleases occurs (Berg et al. 2007).
Most eukaryotes do not maintain canonical plasmids; yeast is a notable exception (SDSU 2007). In addition, the Ti plasmid of the bacterium Agrobacterium tumefaciens can be used to integrate foreign DNA into the genomes of many plants. Other methods of introducing or creating recombinant DNA in eukaryotes include homologous recombination and transfection with modified viruses.
Chimeric plasmids
When recombinant DNA is then further altered or changed to host additional strands of DNA, the molecule formed is referred to as "chimeric" DNA molecule, with reference to the mythological chimera that consisted as a composite of several animals (Berg et al. 2007). The presence of chimeric plasmid molecules is somewhat regular in occurrence, as throughout the lifetime of an organism the propagation by vectors ensures the presence of hundreds of thousands of organismal and bacterial cells that all contain copies of the original chimeric DNA (Berg et al. 2007).
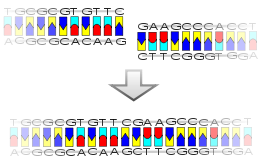
In the production of chimeric plasmids, the processes involved can be somewhat uncertain as the intended outcome of the addition of foreign DNA may not always be achieved and may result in the formation of unusable plasmids (Berg et al. 2007). Initially, the plasmid structure is linearized to allow the addition by bonding of complimentary foreign DNA strands to single-stranded "overhangs" or "sticky ends" present at the ends of the DNA molecule from staggered, or "S shaped" cleavages produced by restriction endonucleases (Berg et al. 2007).
A common vector used for the donation of plasmids originally was the bacterium Escherichia coli and later, the EcoRI derivative, which was used for its versatility with addition of new DNA by "relaxed" replication when inhibited by chloramphenicol and spectinomycin; later being replaced by the pBR322 plasmid (Kaplan et al. 1980). In the case of EcoRI, the plasmid can anneal with the presence of foreign DNA via the route of sticky-end ligation, or with "blunt ends" via blunt-end ligation, in the presence of the phage T4 ligase, which forms covalent links between 3-carbon OH and 5-carbon PO4 groups present on blunt ends (Kaplan et al. 1980). Both sticky-end, or overhang ligation and blunt-end ligation can occur between foreign DNA segments, and cleaved ends of the original plasmid depending upon the restriction endonuclease used for cleavage (Kaplan et al. 1980).
Synthetic insulin production using rDNA
In the 1920s, there was no known way to produce insulin. The problem was solved, temporarily at least, when it was found that insulin from the pancreas of a pig could be used in humans. This method was the primary solution for diabetes until the late twentieth century. The problem with insulin production was raised again: There were not enough pigs to keep the supply of insulin intact. Scientists needed to devise another way. This is where one of the biggest breakthroughs in recombinant DNA technology happened.
The specific gene sequence, or oligonucleotide, that codes for insulin production in humans was introduced to a sample colony of E. coli. Only about 1 out of 106 bacteria picks up the sequence. This is not really a problem though, because the growth rate is only about 30 minutes for E. coli. In a 24 hour period, there may be billions of E. coli that are coded with the DNA sequences needed to induce insulin production (Johnson 1983).
ReferencesISBN links support NWE through referral fees
- Berg, J. M., J. L. Tymoczko, and L. Stryer. 2007. Biochemistry. San Francisco: W. H. Freeman. ISBN 0716787245
- Garret, R. H., and C. M. Grisham. 2000. Biochemistry. Saunders College Publishers. ISBN 0030758173
- Johnson, I. S. 1983. Human insulin from recombinant DNA technology. Science 219(4585): 632-637. Retrieved December 6, 2007.
- Kaplan, N. O., S. P. Colowick, and R. Wu. 1980. Recombinant DNA, Volume 68. Part F (Methods in Enzymology). Academic Press. ISBN 012181968X
- San Diego State University. 2007. Plasmids in eukaryotic microbes: An example. San Diego State University. Retrieved December 6, 2007.
Credits
New World Encyclopedia writers and editors rewrote and completed the Wikipedia article in accordance with New World Encyclopedia standards. This article abides by terms of the Creative Commons CC-by-sa 3.0 License (CC-by-sa), which may be used and disseminated with proper attribution. Credit is due under the terms of this license that can reference both the New World Encyclopedia contributors and the selfless volunteer contributors of the Wikimedia Foundation. To cite this article click here for a list of acceptable citing formats.The history of earlier contributions by wikipedians is accessible to researchers here:
The history of this article since it was imported to New World Encyclopedia:
Note: Some restrictions may apply to use of individual images which are separately licensed.