Cladistics
Cladistics, or phylogenetic systematics, is a system of classifying living and extinct organisms based on evolutionary ancestry as determined by grouping taxa according to "derived characters," that is characteristics or features shared uniquely by the taxa and their common ancestor. Cladistics places heavy emphasis on objective, quantitative analysis and emphasizes evolution and genealogy in contrast to more traditional biological taxonomy with its focus on physical similarities between species. Emphasizing no particular mechanism of evolution, cladistics as a classification schema lies largely separate from much of the debate between those who favor natural selection and those who favor intelligent design.
Cladistics generates diagrams, called "cladograms," that represent the evolutionary tree of life. DNA (deoxyribonucleic acid) and RNA (ribonucleic acid) sequencing data are used in many important cladistic efforts. Cladistics originated in the field of biology by a German entomologist, but in recent years cladistic methods have found application in other disciplines. The word cladistics, created in 1950, is derived from the ancient Greek κλάδος, klados, or "branch."
Although the emphasis of cladistics on biological lineage through millions of years is metaphorically similar to the human convention of tracing genealogical lineage through multiple generations, the two are quite different in substance, as one traces lineage of species while the other traces lineage of specific members of a species. The trend of cladistics toward mapping a connectedness between all species of organisms, based on the theory of descent with modification, shows metaphorical similarity with views of some religions that humans are all connected because of a common origin.
The history of the various schools or research groups that developed around the concept of biological classification was often filled with disputes, competitions, and even bitter opposition (Hull 1988). This is frequently the history of new ideas that challenge the existing paradigm, as cladism has done in presenting a strong alternative to Linnaean taxonomy.
Overview
Systematics is the branch of biology that strives to discover the genealogical relationships underlying organic diversity and also constructs classifications of living things (Sober 1988, 7). There is a diversity of opinion on how genealogy and taxonomy are related.
Two prominent research groups taking very different approaches from each other emerged in the mid-twentieth century (Hull 1988). One, the Sokol-Sneath school, proposed to improve on the methods of traditional Linnaean taxonomy by introducing "numerical taxonomy," which aimed to ascertain the overall similarity among organisms using objective, quantitative, and numerous characters (Hull 1988). A second group, led by the German biologist Willi Hennig (1913-1976), proposed a fundamentally new approach that emphasized classifications representing phylogeny focused on the sister-group relationship: Two taxa are sister groups if they are more related to each other than to a third taxa, and the evidence for this is the presence of characters that the sister groups exhibit but the third group does not exhibit (Hull 1988). That is, the sister groups share a more recent common ancestor with each other than with the third group (Hull 1988). The method emphasizes common ancestry and descent more than chronology. Hennig's 1950 work, Grundzüge einer Theorie der Phylogenetischen Systematik, published in German, began this area of cladistics.
The German-American biologist, Ernst Mayr, in a 1965 paper termed the Sokol-Sneath school "phenetic" because its aim in classifications was to represent the overall similarities exhibited by organisms regardless of descent (Hull 1988). He also coined the term "cladistics" ("branch") for Hennig's system because Hennig wished to represent branching sequences (Hull 1988). Mayr thought his own view to be "evolutionary taxonomy" because it reflected both order of branching (cladistics) and degrees of divergence (phenetics) (Hull 1988).
In Mayr's terms then there would be three notable schools of biological taxonomy: cladists who insist that only genealogy should influence classification; pheneticists who hold that overall similarity, rather than descent, should determine classification; and evolutionary taxonomists (the heirs of traditional Linnaean taxonomists) who hold that both evolutionary descent and adaptive similarity should be used in classification (Sober 1988).
Hennig referred to his approach as phylogenetic systematics, which is the title of his 1966 book. Hennig's major book, even the 1979 version, does not contain the term "cladistics" in the index. A review paper by Dupuis observes that the term clade was introduced in 1958, by Julian Huxley, cladistic by Cain and Harrison in 1960, and cladist (for an adherent of Hennig's school) by Mayr in 1965 (Dupuis 1984). The term "phylogenetics" is often used synonymously with "cladistics."
Computer programs are widely used in cladistics, due to the highly complex nature of cladogram-generation procedures.
Monophyletic groupings
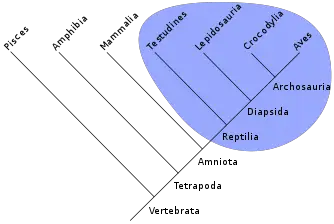
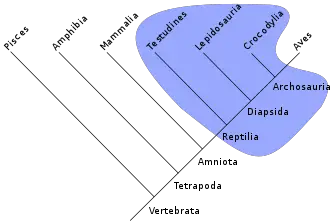
Cladists construct cladograms, branching diagrams, to graphically depict the groups of organisms that share derived characters.
Key to cladistics analysis is identifying monophyletic groups, that is, groups comprising a given species, all of that species' descendants, and nothing else (Sober 1988). In phylogenetics, a group of species is said to be paraphyletic (Greek para meaning near and phyle meaning race) if the group contains its most recent common ancestor, but does not contain all the descendants of that ancestor. For instance, the traditional class Reptilia excludes birds even though they are widely considered to have evolved from an ancestral reptile. Similarly, the traditional invertebrates are paraphyletic because vertebrates are excluded, although the latter evolved from an invertebrate.
A group comprising members from separate evolutionary lines is called polyphyletic. For instance, the once-recognized Pachydermata order was found to be polyphyletic because elephants and rhinoceroses arose separately from non-pachyderms. Evolutionary taxonomists consider polyphyletic groups to be errors in classification, often occurring because convergence or other homoplasy was misinterpreted as homology.
Cladistic taxonomy requires taxa to be clades (monophyletic groups). Cladists argue, therefore, that the prevailing classification system, Linnaean taxonomy, should be reformed to eliminate all non-clades. Others, such as those in the school of evolutionary taxonomy, often use cladistic techniques and require that groups reflect phylogenies, but they also allow both monophyletic and paraphyletic groups as taxa.
Following Hennig, cladists argue that paraphyly is as harmful as polyphyly. The idea is that monophyletic groups can be defined objectively through identifying synapomorphies, that is, features shared uniquely by a group of species and their most immediate common ancestor. This cladistic approach is claimed to be more objective than the alternative approach of defining paraphyletic and polyphyletic groups based on a set of key characteristics determined by researchers. Making such determinations, cladists argue, is an inherently subjective process highly likely to lead to "gradistic" thinking that groups advance from "lowly" grades to "advanced" grades, which can in turn lead to teleological thinking.
Basic procedure
A cladistic analysis organizes a certain set of information by making a distinction between characters and character states. Consider feathers, whose color may be blue in one species but red in another. In this case, "feather-color" is a character and "red feathers" and "blue feathers" are two character states.
In the "old days," before the introduction of computer analysis into cladistics, the researcher would assign the selected character states as being either plesiomorphies, character states present before the last common ancestor of the species group, or synapomorphies, character states that first appeared in the last common ancestor. Usually the researcher would make this assignment by considering one or more outgroups (organisms considered not to be part of the group in question, but nonetheless related to the group). Then, as now, only synapomorphies would be used in characterizing cladistic divisions.
Next, different possible cladograms were drawn up and evaluated by looking for those having the greatest number of synapomorphies. The hope then, as now, was that the number of true synapomorphies in the cladogram would be large enough to overwhelm any unintended symplesiomorphies (homoplasies) caused by convergent evolution, that is, characters that resemble each other because of environmental conditions or function, but not because of common ancestry. A well-known example of homoplasy due to convergent evolution is wings. Though the wings of birds and insects may superficially resemble one another and serve the same function, each evolved independently. If a dataset contained data on a bird and an insect that both scored "POSITIVE" for the character "presence of wings," a homoplasy would be introduced into the dataset, which could cause erroneous results.
When two alternate possible cladograms were evaluated to be equally probable, one was usually chosen based on the principle of parsimony: The most compact arrangement was likely the best hypothesis of relationship (a variation of Occam's razor, which states that the simplest explanation is most often the correct one). Another approach, particularly useful in molecular evolution, involved applying the statistical analysis of maximum likelihood to select the most likely cladogram based on a specific probability model of changes.
Of course, it is no longer done this way: researcher selection, and hence bias, is something to be avoided. These days much of the analysis is done by software: Besides the software to calculate the trees themselves, there is sophisticated statistical software to provide a more objective basis. As DNA sequencing has become easier, phylogenies are increasingly constructed with the aid of molecular data. Computational systematics allows the use of these large data sets to construct objective phylogenies. These can more accurately distinguish some true synapomorphies from homoplasies that are due to parallel evolution. Ideally, morphological, molecular, and possibly other (behavioral, etc.) phylogenies should be combined.
Cladistics does not assume any particular theory of evolution, but it does assume the pattern of descent with modification. Thus, cladistic methods can be, and recently have been, usefully applied to mapping descent with modification in non-biological systems, such as language families in historical linguistics and the filiation of manuscripts in textual criticism.
Cladograms

The starting point of cladistic analysis is a group of species and the molecular, morphological, or other data that characterizes those species. The end result is a tree-like relationship-diagram called a cladogram. The cladogram graphically represents a hypothetical evolutionary process. Cladograms are subject to revision as additional data becomes available.
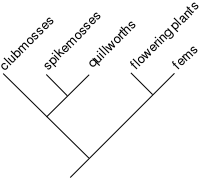
In a cladogram, all organisms lie at the leaves, and each inner node is ideally binary (two-way). The two taxa on either side of a split are called "sister taxa" or "sister groups." Each subtree is called a "clade," and by definition is a natural group, all of whose species share a common ancestor. Each clade is set off by a series of characteristics that appear in its members, but not in the other forms from which it diverged. These identifying characteristics of a clade are its synapomorphies (shared, derived characters). For instance, hardened front wings (elytra) are a synapomorphy of beetles, while circinate vernation, or the unrolling of new fronds, is a synapomorphy of ferns.
Synonyms—The term "evolutionary tree" is often used synonymously with cladogram. The term phylogenetic tree is sometimes used synonymously with cladogram (Singh 2004), but others treat phylogenetic tree as a broader term that includes trees generated with a non-evolutionary emphasis.
Subtrees are clades—In a cladogram, all species lie at the leaves (Albert 2006). The two taxa on either side of a split are called sister taxa or sister groups. Each subtree, whether it contains one item or a hundred thousand items, is called a clade.
Two-way versus three-way Forks—Many cladists require that all forks in a cladogram be 2-way forks. Some cladograms include 3-way or 4-way forks when the data is insufficient to resolve the forking to a higher level of detail, but nodes with more than two branches are discouraged by many cladists.
Depth of a Cladogram—If a cladogram represents N species, the number of levels (the "depth") in the cladogram is on the order of log2(N) (Aldous 1996). For example, if there are 32 species of deer, a cladogram representing deer will be around 5 levels deep (because 25=32). A cladogram representing the complete tree of life, with about 10 million species, would be about 23 levels deep. This formula gives a lower limit: In most cases the actual depth will be a larger value because the various branches of the cladogram will not be uniformly deep. Conversely, the depth may be shallower if forks larger than two-way forks are permitted.
Number of Distinct Cladograms—For a given set of species, the number of distinct rooted cladograms that can in theory be drawn (ignoring which cladogram best matches the species characteristics) is (Lowe 2004):
| Number of Species | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | N |
| Number of Cladograms | 1 | 3 | 15 | 105 | 945 | 10,395 | 135,135 | 2,027,025 | 34,459,425 | 1*3*5*7*...*(2N-3) |
This exponential growth of the number of possible cladograms explains why manual creation of cladograms becomes very difficult when the number of species is large.
Extinct Species in Cladograms—Cladistics makes no distinction between extinct and non-extinct species (Scott-Ram 1990), and it is appropriate to include extinct species in the group of organisms being analyzed. Cladograms based on DNA/RNA generally do not include extinct species because DNA/RNA samples from extinct species are rare. Cladograms based on morphology, especially morphological characteristics preserved in fossils, are more likely to include extinct species.
Time Scale of a Cladogram—A cladogram tree has an implicit time axis (Freeman 1998), with time running forward from the base of the tree to the leaves of the tree. If the approximate date (for example, expressed as millions of years ago) of all the evolutionary forks were known, those dates could be captured in the cladogram. Thus, the time axis of the cladogram could be assigned a time scale (for example 1 cm = 1 million years), and the forks of the tree could be graphically located along the time axis. Such cladograms are called scaled cladograms. Many cladograms are not scaled along the time axis, for a variety of reasons:
- Many cladograms are built from species characteristics that cannot be readily dated (for example, morphological data in the absence of fossils or other dating information)
- When the characteristic data is DNA/RNA sequences, it is feasible to use sequence differences to establish the relative ages of the forks, but converting those ages into actual years requires a significant approximation of the rate of change (Carrol 1997).
- Even when the dating information is available, positioning the cladogram's forks along the time axis in proportion to their dates may cause the cladogram to become difficult to understand or hard to fit within a human-readable format
Summary of terminology
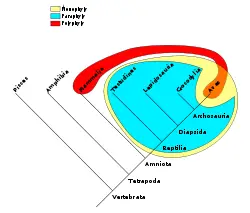
- A clade is an ancestor species and all of its descendants
- A monophyletic group is a clade
- A paraphyletic group is an ancestor species and most of its descendants, usually with a specific group of descendants excluded (for example, reptiles are all the sauropsids (members of the class Sauropsida) except for birds). Most cladists discourage the use of paraphyletic groups.
- A polyphyletic group is a group consisting of members from two non-overlapping monophyletic groups (for example, flying animals). Most cladists discourage the use of polyphyletic groups.
- An outgroup is an organism considered not to be part of the group in question, although it is closely related to the group.
- A characteristic present in both the outgroups and the ancestors is called a plesiomorphy (meaning "close form," as in close to the root ancestor; also called an ancestral state).
- A characteristic that occurs only in later descendants is called an apomorphy (meaning "separate form" or "far from form," as in far from the root ancestor; also called a "derived" state) for that group.
Note: The adjectives plesiomorphic and apomorphic are often used instead of "primitive" and "advanced" to avoid placing value judgments on the evolution of the character states, since both may be advantageous in different circumstances. It is not uncommon to refer informally to a collective set of plesiomorphies as a ground plan for the clade or clades they refer to.
- A species or clade is basal to another clade if it holds more plesiomorphic characters than that other clade. Usually a basal group is very species-poor as compared to a more derived group. It is not a requirement that a basal group be extant. For example, palaeodicots are basal to flowering plants.
- A clade or species located within another clade is said to be nested within that clade.
Cladistics compared with Linnaean taxonomy
Prior to the advent of cladistics, most taxonomists limited themselves to using Linnaean taxonomy for organizing lifeforms. That traditional approach used several fixed levels of a hierarchy, such as Kingdom, Phylum, Class, Order, and Family. Cladistics does not use those terms because one of its fundamental premises is that the evolutionary tree is very deep and very complex, and it is not meaningful to use a fixed number of levels.
Linnaean taxonomy insists that groups reflect phylogenies, but in contrast to cladistics allows both monophyletic and paraphyletic groups as taxa. Since the early twentieth century, Linnaean taxonomists have generally attempted to make genus and lower-level taxa monophyletic.
Cladistics originated in the work of Willi Hennig, and since that time there has been a spirited debate (Wheeler 2000) about the relative merits of cladistics versus Linnaean classification and other Linnaean-associated classification systems, such as the evolutionary taxonomy advocated by Mayr (Benton 2000). Some of the debates that the cladists engaged in had been running since the nineteenth century, but they entered these debates with a new fervor (Hull 1988), as can be learned from the Foreword to Hennig (1979) in which Rosen, Nelson, and Patterson wrote the following—not about Linnaean taxonomy but about the newer evolutionary taxonomy:
Encumbered with vague and slippery ideas about adaptation, fitness, biological species and natural selection, neo-Darwinism (summed up in the "evolutionary" systematics of Mayr and Simpson) not only lacked a definable investigatory method, but came to depend, both for evolutionary interpretation and classification, on consensus or authority (Foreword, page ix).
Proponents of cladistics enumerate key distinctions between cladistics and Linnaean taxonomy as follows (Hennig 1975):
| Cladistics | Linnaean Taxonomy |
| Treats all levels of the tree as equivalent. | Treats each tree level uniquely. Uses special names (such as Family, Class, Order) for each level. |
| Handles arbitrarily-deep trees. | Often must invent new level-names (such as superorder, suborder, infraorder, parvorder, magnorder) to accommodate new discoveries. Biased towards trees about 4 to 12 levels deep. |
| Discourages naming or use of groups that are not monophyletic | Accepts naming and use of paraphyletic groups |
| Primary goal is to reflect actual process of evolution | Primary goal is to group species based on morphological similarities |
| Assumes that the shape of the tree will change frequently, with new discoveries | Often responds to new discoveries by re-naming or re-levelling of Classes, Orders, and Kingdoms |
| Definitions of taxa are objective, hence free from personal interpretation | Definitions of taxa require individuals to make subjective decisions. For example, various taxonomists suggest that the number of Kingdoms is two, three, four, five, or six (see Kingdom). |
| Taxa, once defined, are permanent (e.g. "taxon X comprises the most recent common ancestor of species A and B along with its descendants") | Taxa can be renamed and eliminated (e.g. Insectivora is one of many taxa in the Linnaean system that have been eliminated). |
Proponents of Linnaean taxonomy contend that it has some advantages over cladistics, such as:[1]
| Cladistics | Linnaean Taxonomy |
| Limited to entities related by evolution or ancestry | Supports groupings without reference to evolution or ancestry |
| Does not include a process for naming species | Includes a process for giving unique names to species |
| Difficult to understand the essence of a clade, because clade definitions emphasize ancestry at the expense of meaningful characteristics | Taxa definitions based on tangible characteristics |
| Ignores sensible, clearly-defined paraphyletic groups such as reptiles | Permits clearly-defined groups such as reptiles |
| Difficult to determine if a given species is in a clade or not (for example, if clade X is defined as "most recent common ancestor of A and B along with its descendants," then the only way to determine if species Y is in the clade is to perform a complex evolutionary analysis) | Straightforward process to determine if a given species is in a taxon or not |
| Limited to organisms that evolved by inherited traits; not applicable to organisms that evolved via complex gene-sharing or lateral transfer | Applicable to all organisms, regardless of evolutionary mechanism |
How complex is the Tree of Life?
One of the arguments in favor of cladistics is that it supports arbitrarily complex, arbitrarily deep trees. Especially when extinct species are considered (both known and unknown), the complexity and depth of the tree can be very large. Every single speciation event, including all the species that are now extinct, represents an additional fork on the hypothetical, complete cladogram representing the full tree of life. Fractals can be used to represent this notion of increasing detail: As a viewpoint zooms into the tree of life, the complexity remains virtually constant (Gordon 1999).
This great complexity of the tree and its associated uncertainty is one of the reasons that cladists cite for the attractiveness of cladistics over traditional taxonomy.
Proponents of non-cladistic approaches to taxonomy point to punctuated equilibrium to bolster the case that the tree-of-life has a finite depth and finite complexity. According to punctuated equilibrium, generally a species comes into the fossil record very similar to when it departs the fossil record, as contrasted with phyletic gradualism whereby a species gradually changes over time into another species. If the number of species currently alive is finite, and the number of extinct species that we will ever know about is finite, then the depth and complexity of the tree of life is bounded, and there is no need to handle arbitrarily deep trees.
Applying Cladistics to other disciplines
The processes used to generate cladograms are not limited to the field of biology (Mace 2005).
The generic nature of cladistics means that cladistics can be used to organize groups of items in many different realms. The only requirement is that the items have characteristics that can be identified and measured. For example, one could take a group of 200 spoken languages, measure various characteristics of each language (vocabulary, phonemes, rhythms, accents, dynamics, etc.) and then apply a cladogram algorithm to the data. The result will be a tree that may shed light on how, and in what order, the languages came into existence.
Thus, cladistic methods have recently been usefully applied to non-biological systems, including determining language families in historical linguistics, culture, history (Lipo 2005), and filiation of manuscripts in textual criticism.
Notes
- ↑ Ernst Mayr, Evolution and the Diversity of Life (Selected Essays) (Cambridge, MA: Harvard Univ. Press, 1976). ISBN 0-674-27105-X
ReferencesISBN links support NWE through referral fees
- Albert, V. Parsimony, Phylogeny, and Genomics. Oxford University Press. ISBN 0199297304
- Aldous, D. 1996. Probability distributions on cladograms. In D. J. Aldous, and R. Pemantle, Random Discrete Structures. New York: Springer. ISBN 0387946233
- Ashlock, P. D. 1974. The uses of cladistics. Annual Review of Ecology and Systematics 5: 81-99.
- Benton, M. 2000. Stems, nodes, crown clades, and rank-free lists: Is Linnaeus dead? Biological Reviews 75(4): 633-648.
- Carrol, R. 1997. Patterns and Processes of Vertebrate Evolution. Cambridge University Press. ISBN 052147809X
- Cavalli-Sforza, L. L. and A. W. F. Edwards. 1967. Phylogenetic analysis: Models and estimation procedures. Evol. 21(3): 550-570.
- Cuénot, L. 1940. Remarques sur un essai d'arbre généalogique du règne animal. Comptes Rendus de l'Académie des Sciences de Paris 210: 23-27.
- de Queiroz, K. and J. A. Gauthier. 1992. Phylogenetic taxonomy. Annual Review of Ecology and Systematics 23: 449–480.
- de Queiroz, K. and J. A. Gauthier. 1994. Toward a phylogenetic system of biological nomenclature. Trends in Research in Ecology and Evolution 9(1): 27-31.
- Dupuis, C. 1984. Willi Hennig's impact on taxonomic thought. Annual Review of Ecology and Systematics 15: 1-24.
- Felsenstein, J. 2004. Inferring Phylogenies. Sunderland, MA: Sinauer Associates. ISBN 0878931775
- Freeman, S. 1998. Evolutionary Analysis. Prentice Hall. ISBN 0135680239
- Gordon, R. 1999. The Hierarchical Genome and Differentiation Waves. World Scientific. ISBN 9810222688
- Hamdi, H., H. Nishio, R. Zielinski, and A. Dugaiczyk. 1999. Origin and phylogenetic distribution of Alu DNA repeats: Irreversible events in the evolution of primates. Journal of Molecular Biology 289: 861–871.
- Hennig, W. 1950. Grundzüge einer Theorie der Phylogenetischen Systematik. Berlin: Deutscher Zentralverlag.
- Hennig, W. 1966. Phylogenetic Systematics. Urbana: University of Illinois Press.
- Hennig, W. and W. Hennig. 1982. Phylogenetische Systematik. Berlin: Parey. ISBN 3489609344
- Hennig, W. 1975. Cladistic analysis or cladistic classification: A reply to Ernst Mayr. Systematic Zoology 24: 244-256.
- Hennig, W. 1979. Phylogenetic Systematics. Urbana: University of Illinois Press. ISBN 0252068149
- Hull, D. L. 1979. The limits of cladism. Systematic Zoology 28: 416-440.
- Hull, D. L. 1988. Science as a Process: An Evolutionary Account of the Social and Conceptual Development of Science. Chicago: The University of Chicago Press.
- Kitching, I. J., P. L. Forey, C. J. Humphries, and D. M. Williams. 1998. Cladistics: The Theory and Practice of Parsimony Analysis. Systematics Association Special Volume. 11 (REV): ALL. ISBN 0198501382
- Lipo, C. 2005. Mapping Our Ancestors: Phylogenetic Approaches in Anthropology and Prehistory. Aldine Transaction. ISBN 0202307514
- Lowe, A. 2004. Ecological Genetics: Design, Analysis, and Application. Blackwell Publishing. ISBN 1405100338
- Luria, S., S. J. Gould, and S. Singer. 1981. A View of Life. Menlo Park, CA: Benjamin/Cummings. ISBN 0805366482
- Letunic, I. 2007. Interactive Tree Of Life (iTOL): an online tool for phylogenetic tree display and annotation. Bioinformatics 23(1): 127-128.
- Mace, R. 2005. The Evolution of Cultural Diversity: A Phylogenetic Approach. London: Routledge Cavendish. ISBN 1844720993
- Mayr, E. 1982. The Growth of Biological Thought: Diversity, Evolution and Inheritance. Cambridge, MA: Harvard Univ. Press. ISBN 0674364465
- Mayr, E. 1976. Evolution and the Diversity of Life (Selected essays). Cambridge, MA: Harvard Univ. Press. ISBN 067427105X
- Mayr, E. 1965. Numerical phenetics and taxonomic theory. Systematic Zoology 14:73-97.
- Patterson, C. 1982. Morphological characters and homology. In K. A. Joysey and A. E. Friday, eds., Problems in Phylogenetic Reconstruction. London: Academic Press. ISBN 0123912504
- Rosen, D., G. Nelson, and C. Patterson. 1979. Phylogenetic Systematics. Urbana, IL: University of Illinois Press. ISBN 0252068149
- Scott-Ram, N. R. 1990. Transformed Cladistics, Taxonomy and Evolution. Cambridge University Press. ISBN 0521340861
- Shedlock, A. M., and N. Okada. 2000. SINE insertions: Powerful tools for molecular systematics. Bioessays 22: 148–160.
- Singh, G. 2004. Plant Systematics: An Integrated Approach. Enfield, N.H.: Science. ISBN 1578083516
- Sober, E. 1988. Reconstructing the Past: Parsimony, Evolution, and Inference. Cambridge, MA: The MIT Press. ISBN 026219273X
- Sokal, R. R. 1975. Mayr on cladism—and his critics. Systematic Zoology 24: 257-262.
- Swofford, D. L., G. J. Olsen, P. J. Waddell, and D. M. Hillis. 1996. Phylogenetic inference. In D. M. Hillis, C. Moritz, and B. K. Mable, eds., Molecular Systematics. Sunderland, MA: Sinauer Associates. ISBN 0878932828
- Wheeler, Q. 2000. Species Concepts and Phylogenetic Theory: A Debate. Columbia University Press. ISBN 0231101430
- Wiley, E. O. 1981. Phylogenetics: The Theory and Practice of Phylogenetic Systematics. New York: Wiley Interscience. ISBN 0471059757
- Zwickl, D. J., and D. M. Hillis. 2002. Increased taxon sampling greatly reduces phylogenetic error. Systematic Biology 51: 588-598.
External links
All links retrieved December 10, 2023.
Credits
New World Encyclopedia writers and editors rewrote and completed the Wikipedia article in accordance with New World Encyclopedia standards. This article abides by terms of the Creative Commons CC-by-sa 3.0 License (CC-by-sa), which may be used and disseminated with proper attribution. Credit is due under the terms of this license that can reference both the New World Encyclopedia contributors and the selfless volunteer contributors of the Wikimedia Foundation. To cite this article click here for a list of acceptable citing formats.The history of earlier contributions by wikipedians is accessible to researchers here:
The history of this article since it was imported to New World Encyclopedia:
Note: Some restrictions may apply to use of individual images which are separately licensed.