Actin
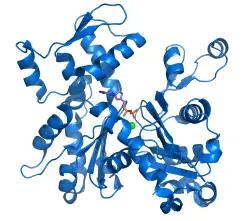
Actin is a globular structural protein that polymerizes in a helical fashion to form an actin filament (or microfilament). These filaments form the cytoskeleton, which is the three-dimensional network inside an eukaryotic cell. The actin filaments provide mechanical support for the cell, determine the cell shape, and enable cell movements through the use of lamellipodia, filopodia, or pseudopodia (cell extensions used for movement). Actin filaments can also participate in certain cell junctions, such as those in cytoplasmic streaming when the cell cytoplasm is flowing, and in contraction of the cell during cytokinesis (division of the cell cytoplasm following nucleus division).
Actin is one of two major muscle proteins that play a crucial role in muscle cell contraction, the other protein being myosin. Actin occurs in two forms, as a monomer and as a polymer. It is as a polymer, F-actin, that it appears as thin filaments, which are interwoven with thick myosin filaments. As these filaments slide past one another, there is a shortening (contraction) and lengthening (relaxation) of the muscle cells.
A relatively simple and ubiquitous protein, being found in most cells, Actin nonetheless demonstrates the extraordinary complexity—and one might say beauty—of creation, requiring a precise order of amino acids folded into a precise three-dimensional shape, and needing myosin, end capping proteins, ATP, and cell signaling mechanisms to function in muscle contraction. And this is but one of innumerable proteins in living organisms.
Actin is one of the most abundant proteins in many eukaryotic cells, with concentrations of over 100 μM. It is also one of the most highly conserved proteins, differing by no more than 5 percent in species as diverse as algae and humans.
Microfilament Assembly
The individual subunits of actin (monomers) are known as globular actin, or for short G-actin. The filamentous polymer composed of individual G-actin subunits is a microfilament and is called F-actin. The microfilaments are the thinnest component of the cytoskeleton, measuring only 7 nanometers in diameter. Much like the microtubules, which are also protein structures found in the cytoskeleton, actin filaments are polar and have two oppositely charged ends. The fast growing plus (+), or barbed, end is opposite from the slow growing minus (-), or pointed, end. The terms barbed and pointed end come from the arrow-like appearance of microfilaments decorated with the motor domain of myosin as seen in electronmicrographs. Filaments elongate approximately ten times faster at the plus (+) end than at the minus (-) end. This phenomenon is known as the treadmill effect.
In the cytosol (fluid component of cytoplasm), actin is predominantly bound to adenosine triphosphate, or ATP. ATP serves as the major source of energy for cellular reactions. Actin can also bind to adenosine diphosphate, or ADP (two phosphate groups). An ATP-actin complex polymerizes faster and dissociates slower than an ADP-actin complex. The process of actin polymerization starts with the association of three G-actin monomers into a trimer. ATP-actin then binds the plus (+) end, and the ATP is subsequently hydrolyzed, which reduces the binding strength between neighboring units and generally destabilizes the filament. ADP-actin dissociates from the minus end and the increase in ADP-actin stimulates the exchange of bound ADP for ATP, leading to more ATP-actin units. This rapid turnover is important for the cell’s movement. End-capping proteins such as CapZ prevent the addition or loss of monomers at the filament end where actin turnover is unfavorable, like in the muscle apparatus.
The protein cofilin binds to ADP-actin units and promotes their dissociation from the minus end and prevents their reassembly. The protein profilin reverses this effect by stimulating the exchange of bound ADP for ATP. In addition, ATP-actin units bound to profilin will dissociate from cofilin and are then free to polymerize. Another important component in filament production is the Arp2/3 complex, which nucleates new actin filaments while bound to existing filaments, thus creating a branched network. All three of these proteins are regulated by cell signaling mechanisms.
Organization
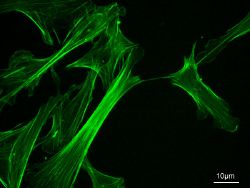
Actin filaments are assembled in two general types of structures: bundles and networks. Actin-binding proteins dictate the formation of either structure since they cross-link actin filaments. Actin filaments have the appearance of a double-stranded helix.
Bundles
In non-muscle actin bundles, the filaments are held together by actin-bundling proteins and/or cationic species so that they are parallel to each other. Bundles play a role in many cellular processes such as cell division (cytokinesis) and cell movement. For example, in vertebrates, the actin-bundling protein villin is almost entirely responsible for causing bundle formations in the microvilli of intestinal cells.
Muscular contraction
Actin, together with myosin filaments, form actomyosin, which provides the mechanism for muscle contraction. Muscular contraction uses ATP for energy. The ATP allows, through hydrolysis, the myosin head to extend up and bind with the actin filament. The myosin head then releases after moving the actin filament in a relaxing or contracting movement by usage of ADP.
In contractile bundles, the actin-bundling protein actinin separates each filament by 40 nm. This increase in distance allows the motor protein myosin to interact with the filament, enabling deformation or contraction. In the first case, one end of myosin is bound to the plasma membrane while the other end walks towards the plus end of the actin filament. This pulls the membrane into a different shape relative to the cell cortex (outer layer of cell). For contraction, the myosin molecule is usually bound to two separate filaments and both ends simultaneously walk towards their filament's plus end, sliding the actin filaments over each other. This results in the shortening, or contraction, of the actin bundle (but not the filament). This mechanism is responsible for muscle contraction and cytokinesis, the division of one cell into two.
Networks
Actin networks, along with many actin-binding proteins (such as the Arp2/3 complex and filamin) form a complex network at the cortical regions of the cell. Recent studies have also suggested that actin network on the cell cortex serve as barriers for molecular diffusion within the plasmic membrane.
Genetics
Actin is one of the most highly conserved proteins, with 80.2 percent gene sequence conservation between Homo sapiens and Saccharomyces cerevisiae (genus of budding fungi) and 95 percent conservation of the primary structure of the protein product. The primary structure refers to the linear sequence of the amino acids in a protein.
Although most yeasts have only a single actin gene, higher eukaryotes generally express several isoforms of actin encoded by a family of related genes. Mammals have at least six actins, which are divided into three classes (alpha, beta and gamma) according to their isoelectric point (the pH at which a substance is electrically zero due to equal concentrations of cations and anions). Alpha actins are generally found in muscle, whereas beta and gamma isoforms are prominent in non-muscle cells. Although there are small differences in sequence and properties between the isoforms, all actins assemble into microfilaments and are essentially identical in the majority of tests performed in vitro, or in an artificial environment outside the living organism.
The typical actin gene has an approximately 100 nucleotide (structural units of RNA and DNA) 5' UTR, a 1200 nucleotide translated region, and a 200 nucleotide 3' UTR. In eukaryotic genetics, the five prime untranslated region (5' UTR) is a particular section of messenger RNA (mRNA) that precedes the translation initiation site of the gene. The three prime untranslated region (3' UTR) consists of the nucleotides present after the stop codon. The majority of actin genes are interrupted by introns, with up to six introns in any of 19 well-characterized locations. Introns are sections of DNA that will be spliced out after transcription, but before the RNA is used. The high conservation of the family makes actin the favored model for studies comparing the introns-early and introns-late models of intron evolution.
All non-spherical prokaryotes appear to possess genes such as MreB that encode homologues of actin; these genes are required for the cell's shape to be maintained. The plasmid-derived gene ParM encodes an actin-like protein whose polymerized form is dynamically unstable and appears to partition the plasmid DNA into the daughter cells during cell division by a mechanism analogous to that employed by microtubules in eukaryotic mitosis.
History
Actin was first observed experimentally in 1887 by W. D. Halliburton, who extracted a protein from muscle that "coagulated" preparations of myosin and which he dubbed "myosin-ferment" (Halliburton 1887). However, Halliburton was unable to further characterize his findings and the discovery of actin is generally credited instead to Brúnó F. Straub, a young biochemist working in Albert Szent-Gyorgyi's laboratory at the Institute of Medical Chemistry at the University of Szeged, Hungary.
In 1942, Straub developed a novel technique for extracting muscle protein that allowed him to isolate substantial amounts of relatively pure actin. Straub's method is essentially the same as that used in laboratories today. Szent-Gyorgyi had previously described the more viscous form of myosin produced by slow muscle extractions as "activated" myosin, and since Straub's protein produced the activating effect, it was dubbed actin. The hostilities of World War II meant that Szent-Gyorgyi and Straub were unable to publish their work in Western scientific journals; it became well-known in the West only in 1945 when information was published as a supplement to the Acta Physiologica Scandinavica (Szent-Gyorgyi 1945).
Straub continued to work on actin and in 1950 reported that actin contains bound ATP and that, during polymerization of the protein into microfilaments, the nucleotide is hydrolyzed to ADP and inorganic phosphate (which remain bound in the microfilament) (Straub 1950). Straub suggested that the transformation of ATP-bound actin to ADP-bound actin played a role in muscular contraction. This, in fact, is only true in smooth muscle and was not experimentally supported until 2001 (Bárány 2001).
The crystal structure of G-actin was solved in 1990 by Kabsch and colleagues (Kabsch 1990). In the same year, a model for F-actin was proposed by Holmes and colleagues (Holmes 1990). The model was derived by fitting a helix of G-actin structures according to low-resolution fiber diffraction data from the filament. Several models of the filament have been proposed since, however, there is still no x-ray structure of F-actin.
ReferencesISBN links support NWE through referral fees
- Bárány, M., J. T. Barron, L. Gu, and K. Bárány. 2001. “Exchange of the actin-bound nucleotide in intact arterial smooth muscle.” J. Biol. Chem. 276: 48398-48403 Entrez PubMed 11602582
- Halliburton, W. D. 1887. “On muscle plasma.” J. Physiol. 8:133.
- Holmes K. C., D. Popp, W. Gebhard, and W. Kabsch. 1990. “Atomic model of the actin filament.” Nature 347: 21-2 Entrez PubMed 2395461
- Kabsch, W., E. G. Mannherz, D. Suck, E. F. Pai, and K. C. Holmes. 1990. “Atomic structure of the actin: DNase I complex.” Nature 347: 37-44 Entrez PubMed 2395459
- Straub, F. B. and G. Feuer. 1950. “Adenosinetriphosphate the functional group of actin.” Biochim. Biophys. Acta. 4: 455-470.
- Szent-Gyorgyi, A. 1945. “Studies on muscle.” Acta Physiol Scandinav 9 (suppl. 25).
Credits
New World Encyclopedia writers and editors rewrote and completed the Wikipedia article in accordance with New World Encyclopedia standards. This article abides by terms of the Creative Commons CC-by-sa 3.0 License (CC-by-sa), which may be used and disseminated with proper attribution. Credit is due under the terms of this license that can reference both the New World Encyclopedia contributors and the selfless volunteer contributors of the Wikimedia Foundation. To cite this article click here for a list of acceptable citing formats.The history of earlier contributions by wikipedians is accessible to researchers here:
The history of this article since it was imported to New World Encyclopedia:
Note: Some restrictions may apply to use of individual images which are separately licensed.
