Pyrimidine
| Pyrimidine | |
|---|---|
| Chemical name | Pyrimidine |
| Chemical formula | C4H4N2 |
| Molecular mass | 80.08796 g/mol |
| CAS number | [289-95-2] |
| Density | 1.016 g/cm3 |
| Melting point | 20–22 °C |
| Boiling point | 123–124 °C |
| SMILES | C1=NC=NC=C1 |
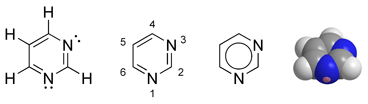
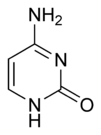
Pyrimidine is a heterocyclic aromatic organic compound similar to benzene and pyridine, containing two nitrogen atoms at positions one and three of the six-member ring (Gilchrist 1997). Heterocyclic compounds are organic compounds (those containing carbon) that contain a ring structure containing atoms in addition to carbon—such as sulfur, oxygen or nitrogen—as part of the ring. Aromaticity is a chemical property in which a conjugated ring of unsaturated bonds, lone pairs, or empty orbitals exhibit a stabilization stronger than would be expected by the stabilization of conjugation alone.
More broadly, pyrimidines is also used in reference to pyrimidine derivatives and structurally related forms.
Pyrimidine is very similar to pyridine, but has an extra nitrogen atom. Pyrimidine is isomeric with two other forms of diazine.
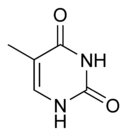
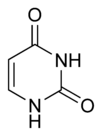
Three nucleobases found in nucleic acids, namely cytosine, thymine, and uracil, are pyrimidine derivatives, with cytosine and thymine in DNA and cytosine and uracil in RNA. All organisms pass on their heredity via the genetic material that is based upon nucleic acids, and every living cell utilizes these pryrimidine nucleobases for transmitting and storing the information needed for manufacturing proteins. The connectedness and unity of all life is thus revealed through the ubiquitousness of these pyrimidine derivatives.
Important pyrimidines (derivatives)
In DNA and RNA, pyrimidines (actually derivatives) cytosine, thymine, and uracil form hydrogen bonds with their complementary purines. In DNA, the purines adenine (A) and guanine (G) pair up with the pyrimidines thymine (T) and cytosine (C) respectively. In RNA, the complement of adenine is uracil instead of thymine and thus the pairs that form are adenine:uracil and guanine and cytosine.
The chemical structures of cytosine, thymine, and uracil are as follows:
These hydrogen bonding modes are for classical Watson-Crick base pairing. Other hydrogen bonding modes ("wobble pairings") are available in both DNA and RNA, although the additional 2'-hydroxyl group of RNA expands the configurations through which RNA can form hydrogen bonds.
Chemical properties
A pyrimidine has many properties in common with pyridine. As the number of nitrogen atoms in the ring increases the ring pi electrons become less energetic and electrophilic aromatic substitution gets more difficult, while nucleophilic aromatic substitution gets easier.
An example of the last reaction type is the displacement of the amino group in 2-aminopyrimidine by chlorine. Reduction in resonance stabilization of pyrimidines may lead to addition and ring cleavage reactions rather than substitutions. One such manifestation is observed in the Dimroth rearrangement.
Compared to pyridine, N-alkylation and N-oxidation are more difficult and pyrimidines are also less basic: the pKa value for protonated pyrimidine is 1.23 compared to 5.30 for pyridine.
Pyrimidine biosynthesis
Synthesis of new pyrimidine
Unlike purines, pyrimidines are assembled before being attached to 5-phosphoribosyl-1-pyrophosphate (PRPP).
The first step begins with formation of carbamoyl phosphate by carbamoyl phosphate synthetase II. This is the regulated step in the pyrimidine biosynthesis.
The second major step is the creation of carbamoyl aspartic acid formed by aspartic transcarbamolyase (aspartate carbamoyl transferase).
The next reaction involves dehydration of the acid catalysed by the enzyme dihhydroorotase to form hydroorotate.
Dihydroorotate then enters the mitochondria where it is oxidized through removal of hydrogens to form orotate. This is the only mitochondrial step in nucleotide rings biosynthesis. The enzyme involved is dihydroorotate dehydrogenase (the only mitochondrial enzyme).
Once orotate is eventually formed, it is combined with PRPP to form orotidine 5' monophosphate (OMP), which is decarboxylated in a reaction catalysed by OMP decarboxylase to form uridine 5' monophosphate (UMP).
UMP is then converted to uridine 5' diphosphate (UDP), catalysed by nucleotide diphosphokinase, which is further phosphorylated to uridine 5' triphosphate (UTP) by CTp synthase. This later reaction eventually leads to the formation of cytidine 5'triphosphate and glutamine is utilized.
Pyrimidine catabolism
Pyrimidines are ultimately catabolized (degraded) to CO2, H2O, and urea. Cytosine can be broken down to uracil, which can be further broken down to N-carbamoyl-β-alanine. Thymine is broken down into β-aminoisobutyrate, which can be further broken down into intermediates eventually leading into the citric acid cycle. β-aminoisobutyrate acts as a rough indicator for rate of DNA turnover.
Organic synthesis
Pyrimidines can also be prepared in the laboratory by organic synthesis. Many methods rely on condensation of carbonyls with amines, for instance the synthesis of 2-Thio-6-methyluracil from thiourea and ethyl acetoacetate (Foster and Snyder 2005), or the synthesis of 4-methylpyrimidine with 4,4-dimethoxy-2-butanone and formamide (Bredereck 2005).
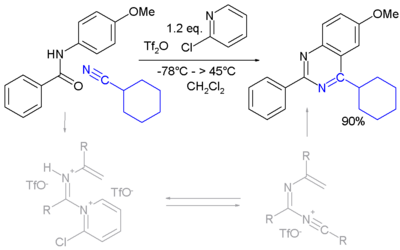
A novel method is by reaction of certain amides with carbonitriles under electrophilic activation of the amide with 2-chloro-pyridine and trifluoromethanesulfonic anhydride (Movassaghi and Hill 2006).
ReferencesISBN links support NWE through referral fees
- Gilchrist, T. L. Heterocyclic Chemistry, 3rd Edition. Essex, England: Addison Wesley, 1997. ISBN 0-582-27843-0
- Movassaghi, M., and M. D. Hill. “Single-step synthesis of pyrimidine derivatives.” J. Am. Chem. Soc. 128(44): 14254-14255, 2006.
| Nucleic acids edit |
|---|
| Nucleobases: Adenine - Thymine - Uracil - Guanine - Cytosine - Purine - Pyrimidine |
| Nucleosides: Adenosine - Uridine - Guanosine - Cytidine - Deoxyadenosine - Thymidine - Deoxyguanosine - Deoxycytidine |
| Nucleotides: AMP - UMP - GMP - CMP - ADP - UDP - GDP - CDP - ATP - UTP - GTP - CTP - cAMP - cGMP |
| Deoxynucleotides: dAMP - dTMP - dUMP - dGMP - dCMP - dADP - dTDP - dUDP - dGDP - dCDP - dATP - dTTP - dUTP - dGTP - dCTP |
| Nucleic acids: DNA - RNA - LNA - PNA - mRNA - ncRNA - miRNA - rRNA - siRNA - tRNA - mtDNA - Oligonucleotide |
Credits
New World Encyclopedia writers and editors rewrote and completed the Wikipedia article in accordance with New World Encyclopedia standards. This article abides by terms of the Creative Commons CC-by-sa 3.0 License (CC-by-sa), which may be used and disseminated with proper attribution. Credit is due under the terms of this license that can reference both the New World Encyclopedia contributors and the selfless volunteer contributors of the Wikimedia Foundation. To cite this article click here for a list of acceptable citing formats.The history of earlier contributions by wikipedians is accessible to researchers here:
The history of this article since it was imported to New World Encyclopedia:
Note: Some restrictions may apply to use of individual images which are separately licensed.