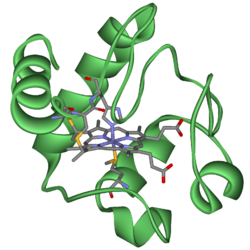
Cytochrome c
Cytochrome c, or cyt c is a small, water soluble heme protein associated with the inner membrane of the mitochondrion. It is an essential link in the electron transport chain through which cells perform the controlled "burning" of glucose and capture much of that released energy by storing it in ATP, the cell's primary energy distribution molecule. Each cytochrome c carries one electron between two different electron transport complexes embedded in the inner membrane. In doing this, cytochrome c repetitively undergoes either oxidation or reduction, but it does not bind oxygen.
Cytochrome c has been particularly thoroughly studied because its small size (about 100 amino acids) and its water solubility permit researchers to isolate it from other mitochondrial proteins, which tend to be not only larger than cytochrome c but also fat soluble and embedded in the membrane. These factors combined have led researchers to determine the amino acid sequences for the cytochrome c occurring in many organisms from yeast to humans.
Cytochrome c is found universally in aerobic organisms, and comparison of amino acid sequences of the molecule in diverse species shows a great deal of similarity among animals, plants, and fungi. Such similarities suggest a common ancestor that must have been using this protein even before basic divergences between plants and animals arose.
Overview
Cytochromes are, in general, membrane-bound hemoproteins that contain heme groups and carry out electron transport. A heme (American English) or haem (British English) is a prosthetic group (the non-protein component of an otherwise protein molecular complex) comprising an iron atom residing in the center of a large heterocyclic organic molecule called a porphyrin. Hemoproteins are part of the larger class of metalloproteins, which includes some complexes whose porphyrin prosthetic group contains at its center a different metal atom than iron.
Cytochromes are found either as monomeric proteins (e.g., cytochrome c) or as subunits of bigger enzymatic complexes that catalyze redox reactions. They are found in both the mitochondrial inner membrane and the endoplasmic reticulum of eukaryotes, in the chloroplasts of plants, in photosynthetic microorganisms, and in bacteria.
The heme group is a highly-conjugated ring system (which means its electrons are very mobile) surrounding an iron ion, which readily interconverts between its primary oxidation states. The iron ion interconverts between the Fe2+ (reduced) and Fe3+ (oxidized) states in electron-transfer processes or between the Fe2+ (reduced) and Fe3+ (formal, oxidized) states in oxidative processes. Cytochromes are, thus, capable of performing oxidation and reduction. Because the cytochromes (as well as other complexes) are held within membranes in an organized way, the redox reactions are carried out in the proper sequence for maximum efficiency.
In the process of oxidative phosphorylation, which is the principal energy-generating process undertaken by organisms that need oxygen to survive, other membrane-bound and -soluble complexes and cofactors are involved in the chain of redox reactions, with the additional net effect that protons (H+) are transported across the mitochondrial inner membrane into the intermembrane space between the inner and outer membranes. The resulting transmembrane proton gradient (protonmotive force) is used to generate ATP, which is the universal chemical energy currency of life. ATP is consumed to drive cellular processes that require energy (such as synthesis of macromolecules, active transport of molecules across the membrane, and assembly of flagella).
Several kinds of cytochromes exist and can be distinguished by spectroscopy, exact structure of the heme group, inhibitor sensitivity, and reduction potential. Three basic types are distinguished by their prosthetic groups: cytochrome a, cytochrome b, and cytochrome d. Cytochrome c, the fourth type of cytochrome, is not defined in terms of the heme group. In mitochondria and chloroplasts, these cytochromes are often combined in electron transport and related metabolic pathways.
Description
Cytochromes c (cytC) are electron-transfer proteins having one or several heme c groups bound to the surrounding protein structure by one or, more generally, two thioether bonds involving sulphydryl groups of cysteine residues. The fifth heme iron ligand is always provided by a histidine residue. Cytochromes c possess a wide range of properties and function in a large number of different redox processes (Pettigrew and Moore 1987).
Cytochrome c's primary structure comprises a chain of 100 amino acids. Its primary function is to transfer electrons between Complexes III and IV in the electron transport chain that drives production of ATP..
R. P. Ambler (1991) recognized four classes of cytochrome c:
- Class I includes the lowspin soluble cytochrome c of mitochondria and bacteria. It has the heme-attachment site toward the N terminus of histidine and the sixth ligand provided by a methionine residue towards the C terminus.
- Class II includes the highspin cytochrome c'. It has the heme-attachment site closed to the N terminus of histidine.
- Class III comprises the low redox potential multiple heme cytochromes. The heme c groups are structurally and functionally nonequivalent and present different redox potentials in the range 0 to -400 mV.
- Class IV was originally created to hold the complex proteins that have other prosthetic groups as well as heme c.
Functions
Role in energy metabolism
Cytochrome c can catalyze several reactions such as hydroxylation and aromatic oxidation, and shows peroxidase activity by oxidation of various electron donors such as 2,2-azino-bis(3-ethylbenzthiazoline-6-sulphonic acid) (ABTS), 2-keto-4-thiomethyl butyric acid and 4-aminoantipyrine.
Role in low level laser therapy
Cytochrome c is also thought to be the functional complex in so called LLLT: Low-level laser therapy. In LLLT, laser light on the wavelength of 670 nanometers penetrates wounded and scarred tissue and increases cellular regeneration. Light of this wavelength appears capable of increasing activity of cytochrome c, thus increasing metabolic activity and freeing up more energy for the cells to repair the tissue.
Role in apoptosis
Cytochrome c is also an intermediate in apoptosis, a controlled form of cell death used to kill cells in the process of development or in response to infection or DNA damage (Liu et al. 1996).
Cytochrome c is released by the mitochondria in response to pro-apoptotic stimuli. The sustained elevation in calcium levels precedes cyt c release from the mitochondria. The release of small amounts of cyt c leads to an interaction with the IP3 receptor (IP3R) on the endoplasmic reticulum (ER), causing ER calcium release. The overall increase in calcium triggers a massive release of cyt c, which then acts in the positive feedback loop to maintain ER calcium release through the IP3Rs. This explains how the ER calcium release can reach cytotoxic levels. This release in turn activates caspase 9, a cysteine protease. Caspase 9 can then go on to activate caspases 3 and 7, which are responsible for destroying the cell from within.
Variation
The cytochrome c molecule of diverse organisms has been studied for the glimpse it gives into evolutionary biology. Cytochrome c is a highly conserved protein across the spectrum of species, found in plants, animals, and many unicellular organisms. This, along with its small size (molecular weight about 12,000 daltons), makes it useful in studies of evolutionary relatedness via cladistics.
The degree of similarities between the cytochrome c from different species correlates closely with the apparent degree of relatedness between species, e.g. the sequences from monkeys and cattle are more similar than the sequences from monkeys and fish. Chickens and turkeys have the identical molecule (amino acid for amino acid) within their mitochondria, whereas ducks possess molecules differing by one amino acid. Similarly, both humans and chimpanzees have the identical molecule, while rhesus monkeys possess cytochromes differing by one amino acid.
ReferencesISBN links support NWE through referral fees
- Ambler, R. P. 1991. Sequence variability in bacterial cytochromes c Biochim. Biophys. Acta 1058(1): 42-47. Retrieved May 16, 2008.
- Bushnell, G. W., G. V. Louie, and G. D. Brayer. 1990. High-resolution three-dimensional structure of horse heart cytochrome c J.Mol.Biol. 214: 585-595. Retrieved May 16, 2008.
- Liu, X., C. Kim, J. Yang, R. Jemmerson, and X. Wang. 1996. Induction of apoptotic program in cell-free extracts: Requirement for dATP and cytochrome c Cell 86(1): 147-57. Retrieved May 16, 2008.
- Pettigrew, G. W. and G. R. Moore. 1987 Cytochromes c: Biological Aspects. New York: SpringerVerlag. ISBN 0387178430.
Further reading
- Broger, C., M. J. Nałecz, and A. Azzi. 1980. Interaction of cytochrome c with cytochrome bc1 complex of the mitochondrial respiratory chain Biochim. Biophys. Acta 592(3): 519-27. PMID 6251869 Retrieved May 16, 2008.
- Evans, M. J., and R. C. Scarpulla. 1989. The human somatic cytochrome c gene: two classes of processed pseudogenes demarcate a period of rapid molecular evolution Proc. Natl. Acad. Sci. U.S.A. 85(24): 9625-9. PMID 2849112 Retrieved May 16, 2008.
- Ferri, K. F., E. Jacotot, J. Blanco, et al. 2001. Mitochondrial control of cell death induced by HIV-1-encoded proteins Ann. N. Y. Acad. Sci. 926: 149-64. PMID 11193032 Retrieved May 16, 2008.
- Garber, E. A., and E. Margoliash. 1990. Interaction of cytochrome c with cytochrome c oxidase: an understanding of the high- to low-affinity transition Biochim. Biophys. Acta 1015(2): 279-87. PMID 2153405 Retrieved May 16, 2008.
- Haider, N. N. Narula, and J. Narula. 2003. Apoptosis in heart failure represents programmed cell survival, not death, of cardiomyocytes and likelihood of reverse remodeling J. Card. Fail. 8(6 Suppl): S512-7. PMID 12555167 Retrieved May 16, 2008.
- Lynch, S. R., D. Sherman, and R. A. Copeland. 1992. Cytochrome c binding affects the conformation of cytochrome a in cytochrome c oxidase J. Biol. Chem. 267(1): 298-302. PMID 1309738 Retrieved May 16, 2008.
- Mannella, C. A. 1998. Conformational changes in the mitochondrial channel protein, VDAC, and their functional implications J. Struct. Biol. 121(2): 207-18. PMID 9615439 Retrieved May 16, 2008.
- Michel, B., and H. R. Bosshard. 1984. Spectroscopic analysis of the interaction between cytochrome c and cytochrome c oxidase J. Biol. Chem. 259(16): 10085-91. PMID 6088481 Retrieved May 16, 2008.
- Skulachev, V. P. 1998. Cytochrome c in the apoptotic and antioxidant cascades FEBS Lett. 423(3): 275-80. PMID 9515723 Retrieved May 16, 2008.
- Tanaka, Y., T. Ashikari, Y. Shibano, et al. 1988. Construction of a human cytochrome c gene and its functional expression in Saccharomyces cerevisiae J. Biochem. 103(6): 954-61. PMID 2844747 Retrieved May 16, 2008.
| ||||||||
Credits
New World Encyclopedia writers and editors rewrote and completed the Wikipedia article in accordance with New World Encyclopedia standards. This article abides by terms of the Creative Commons CC-by-sa 3.0 License (CC-by-sa), which may be used and disseminated with proper attribution. Credit is due under the terms of this license that can reference both the New World Encyclopedia contributors and the selfless volunteer contributors of the Wikimedia Foundation. To cite this article click here for a list of acceptable citing formats.The history of earlier contributions by wikipedians is accessible to researchers here:
The history of this article since it was imported to New World Encyclopedia:
Note: Some restrictions may apply to use of individual images which are separately licensed.