Difference between revisions of "Methionine" - New World Encyclopedia
({{Contracted}}) |
|||
| Line 1: | Line 1: | ||
| − | {{Claimed}} | + | {{Claimed}}{{Contracted}} |
<div> | <div> | ||
<!-- Here is a table of data; skip past it to edit the text. —> | <!-- Here is a table of data; skip past it to edit the text. —> | ||
Revision as of 19:59, 15 June 2007
| Methionine | |
|---|---|
| Systematic name | (S)-2-amino-4-(methylsulfanyl)- butanoic acid |
| Abbreviations | Met M |
| Chemical formula | C5H11NO2S |
| Molecular mass | 149.21 g mol-1 |
| Melting point | 281 °C |
| Density | 1.340 g cm-3 |
| Isoelectric point | 5.74 |
| pKa | 2.16 9.08 |
| CAS number | [63-68-3] |
| PubChem | 876 |
| EINECS number | 200-562-9 |
| SMILES | CSCC[C@H](N)C(O)=O |
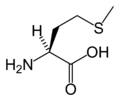  
| |
| Disclaimer and references | |
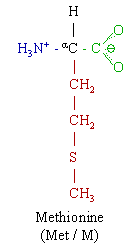
Methionine is an α-amino acid with the chemical formula HO2CCH(NH2)CH2CH2SCH3. This essential is classified as nonpolar. Together with cysteine, methionine is one of two sulfur-containing proteinogenic amino acids. Its derivative S-adenosyl methionine (SAM) serves as a methyl donor. Methionine plays a role in the biosynthesis of cysteine, carnitine, and taurine (by the transsulfuration pathway), lecithin production, the synthesis of phosphatidylcholine, and other phospholipids. Improper conversion of methionine can lead to atherosclerosis.
Methionine is one of only two amino acids encoded by a single codon (AUG) in the standard genetic code (tryptophan, encoded by UGG, is the other). The codon AUG is also significant, in that it carries the "Start" message for a ribosome to begin protein translation from mRNA. As a consequence, methionine is incorporated into the N-terminal position of all proteins in eukaryotes and archaea during translation, although it is usually removed by post-translational modification.
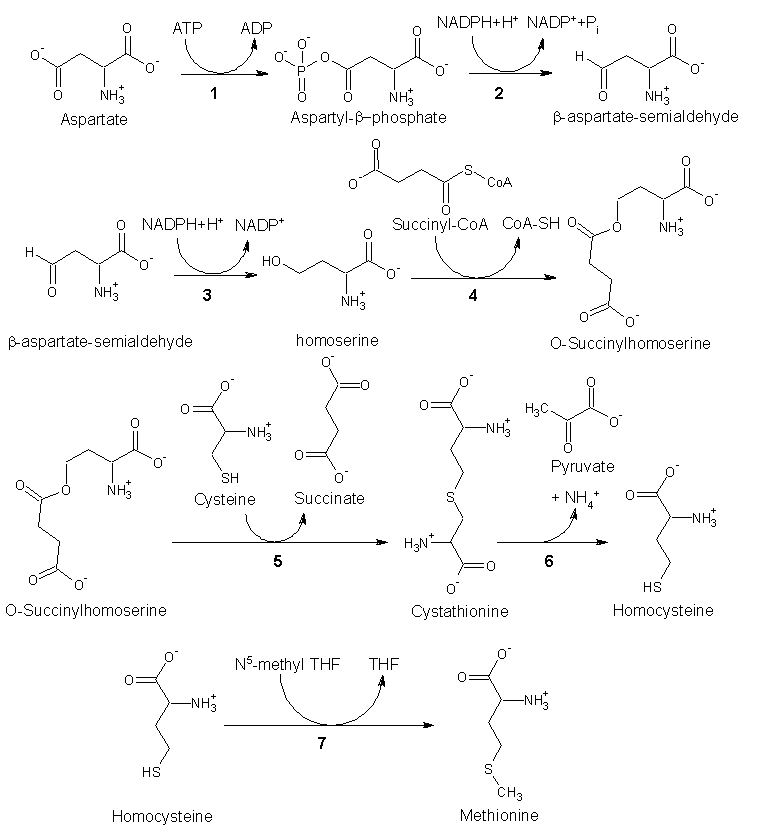
Biosynthesis
As an essential amino acid, methionine is not synthesized in humans, hence we must ingest methionine or methionine-containing proteins. In plants and microorganisms, methionine is synthesized via a pathway that uses both aspartic acid and cysteine. First, aspartic acid is converted via β-aspartyl-semialdehyde into homoserine, introducing the pair of contiguous methylene groups. Homoserine converts to O-succinyl homoserine, which then reacts with cysteine to produce cystathionine, which is cleaved to yield homocysteine. Subsequent methylation of the thiol group by folates affords methionine. Both cystathionine-γ-synthase and cystathionine-β-lyase require Pyridoxyl-5'-phosphate as a cofactor, whereas homocysteine methyltransferase requires Vitamin B12 as a cofactor.[1]
Enzymes involved in methionine biosynthesis:
- aspartokinase
- β-aspartate semialdehyde dehydrogenase
- homoserine dehydrogenase
- homoserine acyltransferase
- cystathionine-γ-synthase
- cystathionine-β-lyase
- methionine synthase (in mammals, this step is performed by homocysteine methyltransferase)
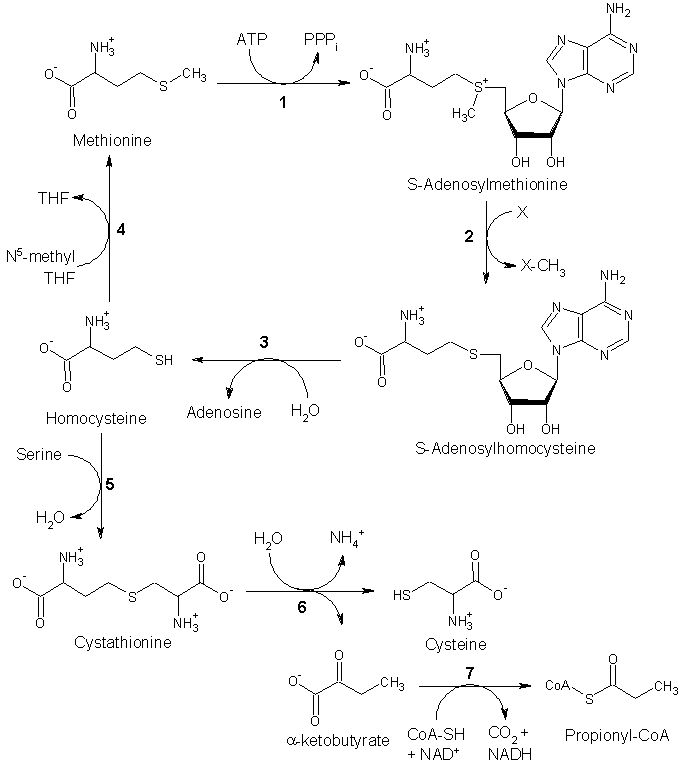
Other biochemical pathways
Although mammals cannot synthesize methionine, they can still utilize it in a variety of biochemical pathways:
Methionine is converted to S-adenosylmethionine (SAM) by (1) methionine adenosyltransferase. SAM serves as a methyl-donor in many (2) methyltransferase reactions and is converted to S-adenosylhomocysteine (SAH). (3) adenosylhomocysteinase converts SAH to homocysteine.
There are two fates of homocysteine.
- First, methionine can be regenerated from homocysteine via (4) methionine synthase. It can also be remethylated using glycine betaine (NNN-trimethyl glycine) to methionine via the enzyme Betaine-homocysteine methyltransferase (E.C.2.1.1.5, BHMT). BHMT makes up to 1.5% of all the soluble protein of the liver, and recent evidence suggests that it may have a greater influence on methionine and homocysteine homeostasis than Methionine sythase.
- Alternatively, homocysteine can be converted to cysteine. (5) cystathionine-β-synthase (a PLP-dependent enzyme) combines homocysteine and serine to produce cystathionine. Instead of degrading cystathionine via cystathionine-β-lyase as in the biosynthetic pathway, cystathionine is broken down to cysteine and α-ketobutyrate via (6) cystathionine-γ-lyase. (7) α-ketoacid dehydrogenase converts α-ketobutyrate to propionyl-CoA, which is metabolized to succinyl-CoA in a three-step process (see propionyl-CoA for pathway).
Synthesis
Racemic methionine can be synthesized from diethyl sodium phthalimidomalonate, (C6H4(CO)2NC(CO2Et)2), by alkylation with chloroethylmethylsulfide, ClCH2CH2SCH3 followed by hydrolysis and decarboxylation.[2]
Dietary aspects
High levels of methionine can be found in sesame seeds, Brazil nuts, fish, meat, and some seeds. Most fruit and vegetables contain very little, although peppers and spinach are the best sources.
See also
- Allantoin
- Formylmethionine
ReferencesISBN links support NWE through referral fees
- ↑ Nelson, D. L.; Cox, M. M. "Lehninger, Principles of Biochemistry" 3rd Ed. Worth Publishing: New York, 2000. ISBN 1-57259-153-6.
- ↑ Barger, G.; Weichselbaum, T. E. “dl-Methionine” Organic Syntheses, Collected Volume 2, p.384 (1943). http://www.orgsyn.org/orgsyn/pdfs/CV2P0384.pdf.
External links
Template:ChemicalSources
| Major families of biochemicals | ||
| Peptides | Amino acids | Nucleic acids | Carbohydrates | Nucleotide sugars | Lipids | Terpenes | Carotenoids | Tetrapyrroles | Enzyme cofactors | Steroids | Flavonoids | Alkaloids | Polyketides | Glycosides | ||
| Analogues of nucleic acids: | The 20 Common Amino Acids | Analogues of nucleic acids: |
| Alanine (dp) | Arginine (dp) | Asparagine (dp) | Aspartic acid (dp) | Cysteine (dp) | Glutamic acid (dp) | Glutamine (dp) | Glycine (dp) | Histidine (dp) | Isoleucine (dp) | Leucine (dp) | Lysine (dp) | Methionine (dp) | Phenylalanine (dp) | Proline (dp) | Serine (dp) | Threonine (dp) | Tryptophan (dp) | Tyrosine (dp) | Valine (dp) | ||
Credits
New World Encyclopedia writers and editors rewrote and completed the Wikipedia article in accordance with New World Encyclopedia standards. This article abides by terms of the Creative Commons CC-by-sa 3.0 License (CC-by-sa), which may be used and disseminated with proper attribution. Credit is due under the terms of this license that can reference both the New World Encyclopedia contributors and the selfless volunteer contributors of the Wikimedia Foundation. To cite this article click here for a list of acceptable citing formats.The history of earlier contributions by wikipedians is accessible to researchers here:
The history of this article since it was imported to New World Encyclopedia:
Note: Some restrictions may apply to use of individual images which are separately licensed.