Salmonella
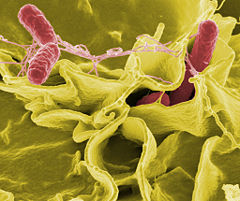
| Salmonella sp. | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
 | ||||||||||||
| Scientific classification | ||||||||||||
| ||||||||||||
|
Salmonella bongori |
Salmonella (plural salmonellae, salmonellas, or salmonella) are any of the various rod-shaped, gram-negative bacteria that comprise the genus Salmonella (family Enterobacteriaceae), some of which are pathogenic. Salmonellosis is the name of a group of infectious diseases caused by salmonella, including typhoid fever, paratyphoid fever, and food poisoning.
Salmonella are found in the intestinal tract of humans, and many animals, including domestic animals, such as chicken and cattle.
Salmonella is a well-known genus because of its ability to cause disease. However, only a few of the more than 2,200 types (serovars or serotypes) of Salmonella cause infections in humans, with the majority of cases traced to only five to ten common forms, mostly S. typhimurium and S. enteritidis (Breslow 2002). Even these infections can be reduced through proper hygiene and personal and social responsibility. Furthermore, salmonella shows promise in the fight against cancer, exhibiting suppression of tumor growth in experimental tests (Nagourney 2001).
Microbiology
Like other members of the bacterial family Enterobacteriaceae, species of Salmonella are gram-negative and rod-shaped. Salmonella do not require oxygen and their main habitat is the intestinal tract of animals. Salmonella species are motile and produce hydrogen sulfide (Giannella et al. 1996). They generally do not ferment lactose.
In a clinical laboratory, Salmonella is usually isolated on MacConkey agar, XLD agar, XLT agar, or DCA agar. Because they cause intestinal infections and are greatly outnumbered by the bacteria normally found in the healthy bowel, primary isolation requires the use of a selective medium, so use of a relatively non-selective medium such as CLED agar is not often practiced. Numbers of salmonella may be so low in clinical samples that stools are routinely also subjected to "enrichment culture" where a small volume of stool is incubated in a selective broth medium, such as selenite broth or Rappaport Vassiliadis soya peptone broth overnight. These media are inhibitory to the growth of the microbes normally found in the healthy human bowel, while allowing salmonellae to become enriched in numbers. Salmonellae may then be recovered by inoculating the enrichment broth on one or more of the primary selective media. On blood agar, they form moist colonies about 2 to 3 millimeters in diameter.
History
Salmonella was named after Daniel Elmer Salmon (1850-1914), an American veterinary pathologist, who described Salmonella enterica (formerly S. choleraesuis). However, it was his colleague and subordinate Theobald Smith (better known for his work on anaphylaxis) who first discovered the bacterium in 1885, from pigs, in an investigation for the cause of hog cholera.
Classification
Salmonella taxonomy is complicated. Tindall et al. (2005) note that "the nomenclature of the genus Salmonella has reached an unsatisfactory state of affairs, with two systems of nomenclature in circulation." One of these systems, proposed in the 1980s by Le Minor and Popoff are widely accepted, but does not conform to the Bacteriological Code, while the other conforms to the rules of the Code but is used by a minority and of decreasing popularity (Tindall et al. 2005). The Judicial Commission of the International Committee for Systematics of Prokaryotes (2005), in Opinion 80, decided that the type species of the genus would be Salmonella enterica and that the type strain would be strain Lt2T. However, Tindall et al. (2005) note that, "like all Opinions, it is limited to matters of nomenclature and does not help to interpret the taxonomic consequences."
As of December 7, 2005, there are two species within the genus Salmonella: Salmonella bongori (previously subspecies V) and Salmonella enterica (formerly called Salmonella choleraesuis), which is divided into six subspecies:
- I—enterica
- II—salamae
- IIIa—arizonae
- IIIb—diarizonae
- IV—houtenae
- V—obsolete (now designated S. bongori)
- VI—indica
There are over 2,200 known serotypes of Salmonella by some accounts (Breslow 2002) and about 4,400 by other accounts (Ryan and Ray 2004). A serovar or serotype is a grouping of microorganisms (or viruses) based on their cell surface antigens, allowing differentiation below the level of species. Serovars may be established based on virulence factors, lipopolysaccharides in gram-negative bacteria, presence of an exotoxin, plasmids, or other characteristics that differentiate two members of the same species (Barron 1996).
The vast majority of human isolates (about 99.5 percent) are subspecies S. enterica. For the sake of simplicity, the Center for Disease Control and Prevention recommends that Salmonella species be referred to only by their genus and serovar, e.g.,
- Salmonella typhi
instead of the more technically correct designation,
- Salmonella enterica subspecies enterica serovar Typhi.
Salmonella isolates are most commonly classified according to serology (Kauffman-White classification) (JCICSP 2005). The main division is first by the somatic O antigen, then by flagellar H antigens. H antigens are further divided into phase 1 and phase 2. The full description of a salmonella isolate is given as (O antigens, Vi : H antigen phase 1: H antigen phase 2).
Note that, with the exception of typhoid fever and paratyphoid, salmonellosis is not a blood-related infection, as is commonly believed.
Examples:
- Salmonella Enteritidis (1,9,12:g,m)
(The O antigens present are 1, 9 and 12; the H antigens are g and m)
- Salmonella Typhi (9,12,Vi:d:−)
(The O antigens are 9, 12,; the H antigen is d: The Vi antigen is associated with the bacterial capsule, which acts as a Virulence factor, hence its name)
In a clinical laboratory, only a small number of serovars are looked for (the remainder being rare or not clinically significant). The Health Protection Agency recommend testing for the following antigens routinely:
- O antigens: 2 4 6.7 8 9 and 3.10
- phase 1 H antigens: 1 2 3 4 5 6 7
- phase 2 H antigens: a b c d E G i r
Isolates that cannot be identified using this panel are sent to the reference laboratory for identification.
Salmonella-associated diseases
Disease-causing Salmonella species have recently been re-classified into a single species, Salmonella enterica, which has numerous serovars. Salmonella Typhi causes typhoid fever. Other salmonellae are frequent causes of foodborne illness, especially from poultry and raw eggs and more generally from food that has been cooked or frozen, and not eaten straight away. Refrigeration does not kill the bacteria, although it can stop their reproduction. While these infections would normally only require a treatment of antibiotics, the overuse of antibiotics in both the poultry and beef industries have created a strain of salmonella that is potentially resistant to antibiotics.
Salmonellosis can also be caught by handling reptiles, such as iguanas or terrapins. A CDC study also noted cases of salmonellosis in 2003 and 2004 associated with handling commercially distributed pet rodents (CDC 2005).
The prevention of salmonellosis as a food illness involves effective sanitizing of food contact surfaces. Alcohol has proven to be an effective topical sanitizer against salmonella. Quaternary ammonium can be used in conjunction with alcohol as a food contact safe sanitizer with increased duration of the sanitizing action. Nonflammable Alcohol Vapor in carbon dioxide NAV-CO2 systems or sodium hypochlorite are frequently used to sanitize surfaces to prevent salmonella.
ReferencesISBN links support NWE through referral fees
- Baron, E. J. 1996. Classification. In S. Baron et al., eds. Baron's Medical Microbiology, 4th edition. University of Texas Medical Branch. ISBN 0963117211
- Breslow, L. 2002. Encyclopedia of Public Health. New York: Macmillan Reference USA/Gale Group Thomson Learning. ISBN 0028658884
- Centers for Disease Control and Prevention. 2005. Outbreak of multidrug-resistant Salmonella Typhimurium associated with rodents purchased at retail pet stores: United States, December 2003-October 2004. Morbidity and Mortality Weekly Report. Retrieved April 9, 2007.
- Giannella, R. A. 1996. Salmonella. In S. Baron et al., eds. Baron's Medical Microbiology, 4th edition. University of Texas Medical Branch. ISBN 0963117211
- Judicial Commission of the International Committee on Systematics of Prokaryotes (JCICSP). 2005. The type species of the genus Salmonella Lignieres 1900 is Salmonella enterica (ex Kauffmann and Edwards 1952) Le Minor and Popoff 1987, with the type strain LT2T, and conservation of the epithet enterica in Salmonella enterica over all earlier epithets that may be applied to this species. Opinion 80. Int J Syst Evol Microbiol 55(Pt 1): 519-520. Retrieved April 9, 2007.
- Nagourney, E. 2001. Vital signs: Treatments; The evil salmonella and its helpful twin. New York Times January 23, 2001. Retrieved April 9, 2007.
- Ryan, K. J., and C. G. Ray (eds). 2004. Sherris Medical Microbiology, 4th ed. McGraw Hill. ISBN 0838585299
- Tindall, B. J., P. A. Grimont, G. H. Garrity, and J. P. Euzéby. 2005. Nomenclature and taxonomy of the genus Salmonella. Int J Syst Evol Microbiol. 55: 521-524. Retrieved April 9, 2007.
External links
All links retrieved December 22, 2022.
- Salmonella Questions and Answers from the Food Safety and Inspection Service of the United States Department of Agriculture.
Credits
New World Encyclopedia writers and editors rewrote and completed the Wikipedia article in accordance with New World Encyclopedia standards. This article abides by terms of the Creative Commons CC-by-sa 3.0 License (CC-by-sa), which may be used and disseminated with proper attribution. Credit is due under the terms of this license that can reference both the New World Encyclopedia contributors and the selfless volunteer contributors of the Wikimedia Foundation. To cite this article click here for a list of acceptable citing formats.The history of earlier contributions by wikipedians is accessible to researchers here:
The history of this article since it was imported to New World Encyclopedia:
Note: Some restrictions may apply to use of individual images which are separately licensed.