Ribosomal RNA
Ribosomal RNA (rRNA) is a type of non-coding ribonucleic acid (RNA) that is a primary and permanent component of ribosomes, the small, cellular particles that form the site of protein synthesis in all living cells. As non-coding RNA, rRNA itself is not translated into a protein, but it does provide a mechanism for decoding messenger RNA (mRNA) into amino acids and interacting with the transfer RNAs (tRNAs) during translation by providing peptidyl transferase activity.
The formation of proteins by rRNA, mRNA, and tRNA is remarkably complex, involving transcription of the various RNAs from DNA, the movement of RNA within a cell, different types of rRNA, and the process of assembling the amino acids in a precise order. And yet this coordinated activity goes on continually in cells, with a single MRNA making several hundred proteins per hour and many thousands of protein molecules per cell generation. With each mammalian cell having millions of ribosomes, and with the human body having many trillions of cells, it is striking to consider how massive, complex, and intricately coordinated is this process of producing proteins for the human body.
Overview
The protein manufacturing unit of all living cells, the ribosome, is composed of ribosomal RNA and protein. It is at the site of the ribosome that messenger RNA's (mRNA) code for linking amino acids together to form new proteins and where transfer RNAs (tRNA) transfer specific amino acids to the growing polypeptide chain during the translation of the mRNA into a protein. The chemical blueprint for the protein product is provided by the mRNA, derived from the DNA genes.
A ribosome can be thought of as a giant enzyme that builds proteins. Its enzymatic activity derives from the presence of the ribosomal RNA (rRNA), which performs the catalytic processes for the synthesis. Meanwhile, the protein portions of the ribosome support the function of the rRNA. More than half the weight of a ribosome is RNA (Alberts et al. 1989).
There are numerous ribosomes in cells—as many as 10 million in a single mammalian cell. Such a cell would need to construct ten million copies of each type of ribosomal RNA molecule. While proteins are rapidly constructed, because each of the many mRNA molecules transcribed from the gene may be translated into as many as 10 protein molecules per minute, and 10,000 protein molecules per mRNA molecule in each cell generation, synthesis of rRNA is not so amplified since these molecules are the final gene product (Alberts et al. 1989). However, adequate rRNA is produced because cells contain multiple copies of the genes that code for rRNA (rRNA genes) (Alberts et al. 1989). E. coli contain seven rRNA genes and human cells contain more than 200 rRNA genes per haploid genome (Alberts et al. 1989).
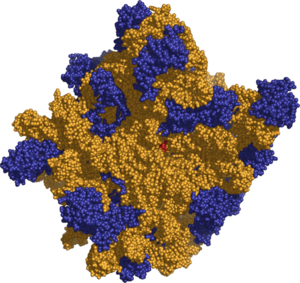
Ribosomes are composed of two subunits, named for how rapidly they sediment when subjected to centrifugation. tRNA is sandwiched between the small and large subunits and the ribosome catalyzes the formation of a peptide bond between the two amino acids that are contained in the tRNA.
A ribosome also has 3 binding sites called A, P, and E.
- The A site in the ribosome binds to an aminoacyl-tRNA (a tRNA bound to an amino acid)
- The NH2 group of the aminoacyl-tRNA which contains the new amino acid attacks the carboxyl group of peptidyl-tRNA (contained within the P site), which contains the last amino acid of the growing chain called peptidyl transferase reaction
- The tRNA that was holding on the last amino acid is moved to the E site, and what used to be the aminoacyl-tRNA is now the peptidyl-tRNA
A single mRNA can be translated simultaneously by multiple ribosomes.
Prokaryote versus eukaryote ribosomes and rRNA
Prokaryote ribosomes are comparatively smaller than eukaryote ribosomes, with a sedimentation coefficient of 70 Svedberg units (abbreviated as 70S), while eukaryote ribosomes have a sedimentation coefficient of 80 Svedberg units (80S).
Both prokaryotic and eukaryotic ribosomes can be broken down into two subunits, with one subunit larger in size and with a dome-like shape and one subunit smaller and located above the larger one, forming a cap-like structure. Each 70S ribosome of prokaryots has a small subunit of 30S and a large subunit of 50S, while each 80S ribosome comprises a small subunit of 40S and a large subunit of 60S. Note that Svedberg measures are not additive because sedimentation rate depends on both mass and surface area.
While the ribosomal subunits are quite similar between prokaryotes and eukaryotes, the 70S ribosomes contain proportionally more RNA than protein, while the 80S ribosomes are composed of less RNA than protein. For example, pea seedlings ribosomes have about 40 percent rRNA and 60 percent protein, while E. coli ribosomes contain 63 percent rRNA and 37 percent protein. In comparing the two subunits themselves, the proportions of rRNA and protein are approximately equal.
The 70S ribosomes have three different types of rRNA: 23S rRNA, 16S rRNA, and 5S r RNA. There are four different types of rRNA in 80s ribosomes: 28s rRNA (but 25-26S rRNA in plants, fungi, and protozoans), 18S rRNA, 5S rRNA, and 5.8S rRNA. These are organized as follows:
| Type | Size | Large subunit | Small subunit |
| prokaryotic | 70S | 50S (5S, 23S) | 30S (16S) |
| eukaryotic | 80S | 60S (5S, 5.8S, 28S) | 40S (18S) |
The 3' end of the 16S rRNA (in a ribosome) binds to a sequence on the 5' end of mRNA called the Shine-Dalgarno sequence.
The 18S rRNA in most eukaryotes is in the small ribosomal subunit, and the large subunit contains three rRNA species (the 5S, 5.8S and 28S rRNAs).
The bacterial 16S, 23S, and 5S rRNA genes are typically organized as a co-transcribed operon. There may be one or more copies of the operon dispersed in the genome, such as the seven of Escherichia coli. Archaea contains either a single rDNA operon or multiple copies of the operon. In contrast, the rRNA genes of eukaryotes generally involves many copies of the genes organized in tandem repeats; for example, in humans, there are approximately 300–400 rDNA repeats present in five clusters (on chromosomes 13, 14, 15, 21, and 22) (Lafontaine and Tollervey 2001).
Mammalian cells have two mitochondrial (12S and 16S) rRNA molecules and four types of cytoplasmic rRNA (28S, 5.8S, 5S (large ribosome subunit) and 18S (small subunit). The 28S, 5.8S, and 18S rRNAs are encoded by a single transcription unit (45S) separated by two internally transcribed spacers (ITS). The 45S rDNA is organized into 5 clusters (each has 30-40 repeats) on chromosomes 13, 14, 15, 21, and 22. These are transcribed by RNA polymerase I. 5S occurs in tandem arrays (~200-300 true 5S genes and many dispersed pseudogenes), the largest one on the chromosome 1q41-42. 5S rRNA is transcribed by RNA polymerase III.
The tertiary structure of the small subunit ribosomal RNA (SSU rRNA) has been resolved by X-ray crystallography (Yusupov et al. 2001). The secondary structure of SSU rRNA contains 4 distinct domains—the 5', central, 3' major and 3' minor domains. A model of the secondary structure for the 5' domain (500-800 nucleotides) is shown.
Translation
Translation is the net effect of proteins being synthesized by ribosomes, from a copy (mRNA) of the DNA template in the nucleus. One of the components of the ribosome (16s rRNA) base pairs complementary to a sequence upstream of the start codon in mRNA.
Importance of rRNA
In addition to their enzymatic role in the synthesis of proteins, ribosomal RNA has important applications in medicine and in evolutionary biology.
In medicine, the difference between prokaryote and eukaryote ribosomes are exploited to create antibiotics to destory a bacterial infection without damaging the cells of an infected person. For example, bacterial 70S ribosomes are vulnerable to chloramphenicol, while the eukaryotic 80S ribosomes are not vulnerable. Ribosomal RNA is the target of such clinically relevant antibiotics as erythromycin, kasugamycin, micrococcin, paromomycin, chloramphenicol, spectinomycin, streptomycin, and thiostrepton.
In evolutionary biology, ribosomal RNA is considered the most conserved (least variable) gene in all cells (Smit et al. 2007). (The proteins in ribosomes have been poorly conserved (Alberts et al. 1989).) For this reason, genes that encode the rRNA (rDNA) are sequenced to identify an organism's taxonomic group, calculate related groups, and estimate rates of species divergence. As a result, many thousands of rRNA sequences are known and stored in specialized databases such as RDP-II (Cole et al. 2003) and the European SSU database (Wuyts et al. 2002).
ReferencesISBN links support NWE through referral fees
- Alberts, B., D. Bray, J. Lewis, M. Raff, K. Roberts, and J. D. Watson. Molecular Biology of the Cell, 2nd edition. New York: Garland Publishing, 1989. ISBN 0824036956.
- Alberts, B., A. Johnson, J. Lewis, M. Raff, K. Roberts, and P. Walter. 2002. Molecular Biology of the Cell, 4th edition. New York: Garland Science. ISBN 0815332181.
- Cole, J. R., B. Chai, T. L. Marsh, R. J. Farris, Q. Wang, S. A. Kulam, S. Chandra, D. M. McGarrell, T. M. Schmidt, G. M. Garrity, and J. M. Tiedje. 2003. The Ribosomal Database Project (RDP-II): Previewing a new autoaligner that allows regular updates and the new prokaryotic taxonomy. Nucleic Acids Res 31: 442-443. PMID 12520046. Retrieved October 4, 2008.
- Lafontaine, D. L. J., and D. Tollervey. 2001. Ribosomal RNA. Encyclopedia of Life Sciences. Retrieved October 4, 2008.
- Smit, S., J. Widmann, and R. Knight. 2007. Evolutionary rates vary among rRNA structural elements. Nucleic Acids Res 35(10): 3339–3354. PMID 17468501. Retrieved October 4, 2008.
- Wuyts, J., Y. Van de Peer, T. Winkelmans, and R. De Wachter. 2002. The European database on small subunit ribosomal RNA. Nucleic Acids Res 30: 183-185. PMID 11752288. Retrieved October 4, 2008.
- Yusupov, M. M., G. Z. Yusupova, A. Baucom, et al. 2001. Crystal structure of the ribosome at 5.5 A resolution. Science 292(5518): 883–896. PMID 11283358. Retrieved October 4, 2008.
External links
All links retrieved December 8, 2022.
- SILVA rRNA Database Project (also includes Eukaryotes (18S) and LSU (23S/28S))
- European database on small subunit ribosomal RNA
- Ribosomal Database Project II
Credits
New World Encyclopedia writers and editors rewrote and completed the Wikipedia article in accordance with New World Encyclopedia standards. This article abides by terms of the Creative Commons CC-by-sa 3.0 License (CC-by-sa), which may be used and disseminated with proper attribution. Credit is due under the terms of this license that can reference both the New World Encyclopedia contributors and the selfless volunteer contributors of the Wikimedia Foundation. To cite this article click here for a list of acceptable citing formats.The history of earlier contributions by wikipedians is accessible to researchers here:
The history of this article since it was imported to New World Encyclopedia:
Note: Some restrictions may apply to use of individual images which are separately licensed.