Transcription (genetics)
This article is part of the series on: Gene expression | |||
| Introduction to Genetics | |||
| General flow: DNA > RNA > Protein | |||
| special transfers (RNA > RNA, RNA > DNA, Protein > Protein) | |||
| Genetic code | |||
| Transcription | |||
| Transcription (Transcription factors, RNA Polymerase,promoter) | |||
| post-transcriptional modification (hnRNA,Splicing) | |||
| Translation | |||
| Translation (Ribosome,tRNA) | |||
| post-translational modification (functional groups, peptides, structural changes) | |||
| gene regulation | |||
| epigenetic regulation (Hox genes, Genomic imprinting) | |||
| transcriptional regulation | |||
| post-transcriptional regulation (sequestration, alternative splicing,miRNA) | |||
| post-translational regulation (reversible,irrevesible) | |||
In biology, transcription is the cellular process of synthesizing RNA based on a DNA template. DNA transcription generates the information-carrying messenger RNAs (mRNAs) used for protein synthesis as well as the other RNA molecules (transfer RNA, ribosomal RNA, etc.) that have catalytic and structural roles in the cell.
In transcription, molecules of RNA are synthesized based on the information stored in DNA, although utilizing only a portion of the DNA molecule to produce the much smaller RNAs. Both nucleic acid sequences, DNA and RNA, use complementary language, and the information is simply transcribed, or copied, from one molecule to the other. One significant difference between the RNA and DNA sequences is the substitution of the base uracil (U) in RNA in place of the closely related base thymine (T) of DNA. Both of these bases pair with adenine (A).
The process of transcription, which is critical for all life and serves as the first stage in building proteins, is very complex and yet remarkably precise. The harmony underlying nature is reflected in the intricate coordination involved in producing RNA molecules from particular segments of the DNA molecule.
Overview of basic process
Transcription, or RNA synthesis, is the process of transcribing DNA nucleotide sequence information into RNA sequence information. The RNA retains the information of the specific region of the DNA sequence from which it was copied.
DNA transcription is similar to DNA replication in that one of the two strands of DNA acts as a template for the new molecule. However, in DNA replication, the new strand formed remains annealed to the DNA strand from which it was copied, whereas in DNA transcription the single-stranded RNA product does not remain attached to the DNA strand, but rather is released as the DNA strand reforms. In addition, RNA molecules are short and are only copied from a portion of the DNA (Alberts et al. 1989).
Transcription has some proofreading mechanisms, but they are fewer and less effective than the controls for copying DNA; therefore, transcription has a lower copying fidelity than DNA replication (Berg et al. 2006).
Synthesis of RNA molecules is done by RNA polymerase enzymes. Eukaryotes have different RNA polymerase molecules to synthesize different types of RNA but most of our knowledge of RNA polymerase comes from the single enzyme that mediates all RNA synthesis in bacteria (Alberts et al. 1989). Both bacterial and eukaryotic RNA polymerases are large, complicated molecules with a total mass of over 500,000 daltons (Alberts et al. 1989).
The stretch of DNA that is transcribed into an RNA molecule is called a transcription unit. A DNA transcription unit that is translated into protein contains sequences that direct and regulate protein synthesis in addition to coding the sequence that is translated into protein. RNA molecules, like DNA molecules, have directionality, which is indicated by reference to either the 5’ end or the 3’ (three prime) end (Zengel 2003). The regulatory sequence that is before (upstream (-), towards the 5' DNA end) the coding sequence is called 5' untranslated region (5'UTR), and sequence found following (downstream (+), towards the 3' DNA end) the coding sequence is called 3' untranslated region (3'UTR).
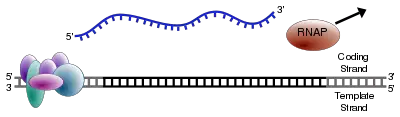
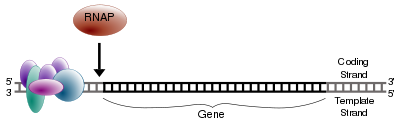
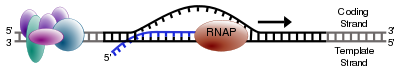
As in DNA replication, RNA is synthesized in the 5' → 3' direction (from the point of view of the growing RNA transcript). Only one of the two DNA strands is transcribed. This strand is called the “template strand,” because it provides the template for ordering the sequence of nucleotides in an RNA transcript. The other strand is called the coding strand, because its sequence is the same as the newly created RNA transcript (except for uracil being substituted for thymine). The DNA template strand is read 3' → 5' by RNA polymerase and the new RNA strand is synthesized in the 5'→ 3' direction.
The RNA polymerase enzyme begins synthesis at a specific start signal on the DNA (called a promoter) and ends its synthesis at a termination signal, whereupon the complete RNA chain and the polymerase are released (Alberts et al. 1989). Essentially, a polymerase binds to the 3' end of a gene (promoter) on the DNA template strand and travels toward the 5' end. The promoter determines which of the two strands of DNA are transcribed for the particular region of DNA being transcribed (Alberts et al. 1989). During transcription, the RNA polymerase, after binding to promoter, opens up a region of DNA to expose the nucleotides and moves stepwise along the DNA, unwinding the DNA to expose areas for transcription, and ends when it encounters the termination signal (Alberts et al. 1989).
One function of DNA transcription is to produce messenger RNAs for the production of proteins via the process of translation. DNA sequence is enzymatically copied by RNA polymerase to produce a complementary nucleotide RNA strand, called messenger RNA (mRNA), because it carries a genetic message from the DNA to the protein-synthesizing machinery of the cell in the ribosomes. In the case of protein-encoding DNA, transcription is the first step that usually leads to the expression of the genes, by the production of the mRNA intermediate, which is a faithful transcript of the gene's protein-building instruction.
In mRNA, as in DNA, genetic information is encoded in the sequence of four nucleotides arranged into codons of three bases each. Each codon encodes for a specific amino acid, except the stop codons that terminate protein synthesis. With four different nucleotides, there are 64 different codons possible. All but three of these combinations (UAA, UGA, and UAG—the stop codons) code for a particular amino acid. However, there are only twenty amino acids, so some amino acids are specified by more than one codon (Zengel 2003).
Unlike DNA replication, mRNA transcription can involve multiple RNA polymerases on a single DNA template and multiple rounds of transcription (amplification of particular mRNA), so many mRNA molecules can be produced from a single copy of a gene.
DNA transcription also produces transfer RNAs (tRNAs), which also are important in protein synthesis. Transfer RNAs transport amino acids to the ribosomes and then act to transfer the correct amino acid to the correct part of the growing polypeptide. Transfer RNAs are small noncoding RNA chains (74-93 nucleotides). They have a site for amino acid attachment, and a site called an anticodon. The anticodon is an RNA triplet complementary to the mRNA triplet that codes for their cargo amino acid. Each tRNA transports only one particular amino acid.
Transcription is divided into 5 stages: Pre-initiation, initiation promoter clearance, elongation, and termination.
Prokaryotic vs. eukaryotic transcription
There are a number of significant differences between prokaryotic transcription and eukaryotic transcription.
A major distinction is that prokaryotic transcription occurs in the cytoplasm alongside translation. Eukaryotic transcription is localized to the nucleus, where it is separated from the cytoplasm by the nuclear membrane. The transcript is then transported into the cytoplasm where translation occurs.
Another important difference is that eukaryotic DNA is wound around histones to form nucleosomes and packaged as chromatin. Chromatin has a strong influence on the accessibility of the DNA to transcription factors and the transcriptional machinery including RNA polymerase.
In prokaryotes, mRNA is not modified. Eukaryotic mRNA is modified through RNA splicing, 5' end capping, and the addition of a polyA tail.
All RNA synthesis is mediated by a single RNA polymerase molecule, while in eukaryotes there are three different RNA polymerases, one making all of the mRNAs for protein synthesis and the others making RNAs with structural and catalytic roles (tRNAs, rRNAs, and so on)
Pre-Initiation
Unlike DNA replication, transcription does not need a primer to start. RNA polymerase simply binds to the DNA and, along with other co-factors, unwinds the DNA to create an initial access to the single-stranded DNA template. However, RNA Polymerase does require a promoter, like the ation bubble, so that the RNA polymerase has sequence.
Proximal (core) Promoters: TATA promoters are found around -10 and -35 bp to the start site of transcription. Not all genes have TATA box promoters and there exists TATA-less promoters as well. The TATA promoter consensus sequence is TATA(A/T)A(A/T). Some strong promoters have UP sequences involved so that the certain RNA polymerases can bind in greater frequencies.
The following are the steps involved in TATA Promoter Complex formation: 1. General transcription factors bind 2. TFIID, TFIIA, TFIIB, TFIIF (w/RNA Polymerase), TFIIH/E The complex is called the closed pre-initiation complex and is closed. Once the structure is opened by TFIIH initiation starts.
Initiation
In bacteria, transcription begins with the binding of RNA polymerase to the promoter in DNA. The RNA polymerase is a core enzyme consisting of five subunits: 2 α subunits, 1 β subunit, 1 β' subunit, and 1 ω subunit. At the start of initiation, the core enzyme is associated with a sigma factor (number 70) that aids in finding the appropriate -35 and -10 basepairs downstream of promoter sequences.
Transcription initiation is far more complex in eukaryotes, the main difference being that eukaryotic polymerases do not directly recognize their core promoter sequences. In eukaryotes, a collection of proteins called transcription factors mediate the binding of RNA polymerase and the initiation of transcription. Only after certain transcription factors are attached to the promoter does the RNA polymerase bind to it. The completed assembly of transcription factors and RNA polymerase bind to the promoter, called transcription initiation complex. Transcription in archaea is similar to transcription in eukaryotes (Quhammouch et al. 2003).
Promoter clearance
After the first bond is synthesized, the RNA polymerase must clear the promoter. During this time there is a tendency to release the RNA transcript and produce truncated transcripts. This is called abortive initiation and is common for both eukaryotes and prokaroytes. Once the transcript reaches approximately 23 nucleotides it no longer slips and elongation can occur. This is an ATP dependent process.
Promoter clearance also coincides with phosphorylation of serine 5 on the carboxy terminal domain which is phosphorylated by TFIIH.
Elongation
One strand of DNA, the template strand (or coding strand), is used as a template for RNA synthesis. As transcription proceeds, RNA polymerase traverses the template strand and uses base pairing complementarity with the DNA template to create an RNA copy. Although RNA polymerase traverses the template strand from 3' → 5', the coding (non-template) strand is usually used as the reference point, so transcription is said to go from 5' → 3'. This produces an RNA molecule from 5' → 3', an exact copy of the coding strand (except that thymines are replaced with uracils, and the nucleotides are composed of a ribose (5-carbon) sugar where DNA has deoxyribose (one less oxygen atom) in its sugar-phosphate backbone).
In producing mRNA, multiple RNA polymerases can be involved on a single DNA template and result in many mRNA molecules from a single gene via multiple rounds of transcription.
This step also involves a proofreading mechanism that can replace incorrectly incorporated bases.
Prokaryotic elongation starts with the "abortive initiation cycle." During this cycle RNA polymerase will synthesize mRNA fragments 2-12 nucleotides long. This continues to occur until the σ factor rearranges, which results in the transcription elongation complex (which gives a 35 bp moving footprint). The σ factor is released before 80 nucleotides of mRNA are synthesized.
In eukaryotic transcription, the polymerase can experience pauses. These pauses may be intrinsic to the RNA polymerase or due to chromatin structure. Often the polymerase pauses to allow appropriate RNA editing factors to bind.
Termination
Bacteria use two different strategies for transcription termination. In Rho-independent transcription termination, RNA transcription stops when the newly synthesized RNA molecule forms a G-C rich hairpin loop, followed by a run of U's, which makes it detach from the DNA template. In the "Rho-dependent" type of termination, a protein factor called "Rho" destabilizes the interaction between the template and the mRNA, thus releasing the newly synthesized mRNA from the elongation complex.
Transcription termination in eukaryotes is less well understood. It involves cleavage of the new transcript, followed by template-independent addition of As at its new 3' end, in a process called polyadenylation.
Transcription factories
Active transcription units are clustered in the nucleus, in discrete sites called “transcription factories.” Such sites could be visualized after allowing engaged polymerases to extend their transcripts in tagged precursors (Br-UTP or Br-U), and immuno-labeling the tagged nascent RNA. Transcription factories can also be localized using fluorescence in situ hybridization, or marked by antibodies directed against polymerases. There are ~10,000 factories in the nucleoplasm of a HeLa cell, among which are ~8,000 polymerase II factories and ~2,000 polymerase III factories. Each polymerase II factor contains ~8 polymerases. As most active transcription units are associated with only one polymerase, each factory will be associated with ~8 different transcription units. These units might be associated through promoters and/or enhancers, with loops forming a "cloud" around the factor.
History
A molecule that allows the genetic material to be realized as a protein was first hypothesized by Jacob and Monod. RNA synthesis by RNA polymerase was established in vitro by several laboratories by 1965; however, the RNA synthesized by these enzymes had properties that suggested the existence of an additional factor needed to terminate transcription correctly.
In 1972, Walter Fiers became the first person to actually prove the existence of the terminating enzyme.
Roger D. Kornberg won the 2006 Nobel Prize in Chemistry "for his studies of the molecular basis of eukaryotic transcription" (NF 2006).
Reverse transcription
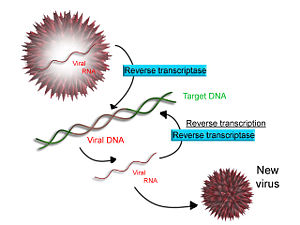
Some viruses (such as HIV), have the ability to transcribe RNA into DNA. HIV has an RNA genome that is duplicated into DNA. The resulting DNA can be merged with the DNA genome of the host cell.
The main enzyme responsible for synthesis of DNA from an RNA template is called reverse transcriptase. In the case of HIV, reverse transcriptase is responsible for synthesizing a complementary DNA strand (cDNA) to the viral RNA genome. An associated enzyme, ribonuclease H, digests the RNA strand, and reverse transcriptase synthesizes a complementary strand of DNA to form a double helix DNA structure. This cDNA is integrated into the host cell's genome via another enzyme (integrase) causing the host cell to generate viral proteins, which reassemble into new viral particles. Subsequently, the host cell undergoes programmed cell death (apoptosis).
Some eukaryotic cells contain an enzyme with reverse transcription activity called telomerase. Telomerase is a reverse transcriptase that lengthens the ends of linear chromosomes. Telomerase carries an RNA template from which it synthesizes DNA repeating sequence, or "junk" DNA. This repeated sequence of "junk" DNA is important because every time a linear chromosome is duplicated, it is shortened in length. With "junk" DNA at the ends of chromosomes, the shortening eliminates some repeated, or junk sequence, rather than the protein-encoding DNA sequence that is further away from the chromosome ends. Telomerase is often activated in cancer cells to enable cancer cells to duplicate their genomes without losing important protein-coding DNA sequence. Activation of telomerase could be part of the process that allows cancer cells to become technically immortal.
ReferencesISBN links support NWE through referral fees
- Alberts, B., D. Bray, J. Lewis, M. Raff, K. Roberts, and J. D. Watson. 1989. Molecular Biology of the Cell, 2nd edition. New York: Garland Publishing. ISBN 0824036956.
- Berg, J., J. L. Tymoczko, and L. Stryer. 2006. Biochemistry, 6th edition. San Francisco: W. H. Freeman. ISBN 0716787245.
- Brooker, R. J. 2005. Genetics: Analysis and Principles, 2nd edition. New York: McGraw-Hill.
- Ouhammouch, M., R. E. Dewhurst, W. Hausner, M. Thomm, and E. P. Geiduschek. 2003. Activation of archaeal transcription by recruitment of the TATA-binding protein. Proceedings of the National Academy of Sciences of the United States of America 100(9): 5097–5102. PMID 12692306. Retrieved February 20, 2009.
- Nobel Foundation (NF). 2006. The Nobel Prize in Chemistry 2006: Roger D. Kornberg. Nobel Foundation. Retrieved February 20, 2009.
- Zengel, J. 2003. Translation. In R. Robinson, Genetics. New York: Macmillan Reference USA. OCLC 55983868.
| |||||||||||
| |||||
Credits
New World Encyclopedia writers and editors rewrote and completed the Wikipedia article in accordance with New World Encyclopedia standards. This article abides by terms of the Creative Commons CC-by-sa 3.0 License (CC-by-sa), which may be used and disseminated with proper attribution. Credit is due under the terms of this license that can reference both the New World Encyclopedia contributors and the selfless volunteer contributors of the Wikimedia Foundation. To cite this article click here for a list of acceptable citing formats.The history of earlier contributions by wikipedians is accessible to researchers here:
The history of this article since it was imported to New World Encyclopedia:
Note: Some restrictions may apply to use of individual images which are separately licensed.