Genetic recombination
Genetic recombination is one of several mechanisms leading to the variation observed in the genetic inheritance passed down the generations by sexual reproduction. More technically put, genetic recombination is the transmission-genetic process by which the combinations of alleles—any of a number of viable DNA coding of the same gene observed at different loci (part of a chromosome)—in two parental individuals become shuffled in offspring individuals. This definition is commonly used in classical transmission genetics, evolutionary biology, and population genetics.
Such shuffling can be the result of recombination via intra-chromososomal recombination or “crossing over,” and via inter-chromososomal recombination also called independent assortment. Recombination therefore only shuffles already existing genetic variation and does not create new variation at the involved loci.
Genetic recombination underlies the changes that occur during evolution. In evolutionary biology, genetic recombination, be it inter- or intra-chromososomal, is thought to have many advantages. The simplest advantage is the avoidance of DNA degradation or "Muller's ratchet." Muller's ratchet is the name given to the process by which the genomes of an asexual population accumulate deleterious mutations in an irreversible manner (hence the word ratchet)—a process that the genomes of sexual populations can easily reverse thanks to recombination.
As a result of recombination, individuals are produced with novel assortments of genes, adding to the reality that individuals produced by sexual reproduction usually differ genetically from each other and from both parents. However, genetic recombination remains poorly understood.
Crossing over
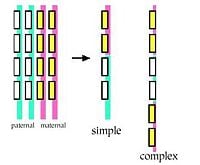
The least complicated, though not well understood, example of sexual reordering of DNA is the basic switching of maternal and paternal DNA. Such crossing over of two chromosomes inherited from one's parents occurs during meiosis. Simply put, genetic information is carried on the chromosomes, with one set inherited from the father, one set inherited from the mother. When a pair of chromosomes "recombine" during sex cell formation, in the simplest case, genetic information on the paternal chromosome is switched with information on the maternal one, and vice versa. The two 'daughter' chromosomes that result are not identical to either of the original parental ones.
To explain further, the sexual reproductive cycle involves the alteration of diploid cells carrying a double set of chromosomes with haploid cells involving a single set of chromosomes. The diploid germ cell, which divides into the haploid sex cells, has two copies of the genome, with one set of each chromosome from the male parent and one set of each chromosome from the female parent—each as a double helix, so there are four strands of DNA involved. Through the process of meiosis, the diploid germ cell produces haploid sex cells such as egg or sperm that have one copy of the genome or one set of chromosomes as a single double helix. It should be noted that during meiosis, there is duplication (DNA replication) and pairing of chromosomes prior to cell division to form haploid cells. To complete the reproductive cycle, haploid cells from a male parent and a female parent fuse at fertilization to form new diploid cells.
The process of genetic recombination or crossing over occurs at the stage of the sex-cell creation (meiosis), when there are four copies of the genome present (involving eight strands of DNA), all aligned with one another on the spindle axis.
This tetraplex or tetrad state (where the eight DNA strands of the four copies of each chromosome are being reordered) lasts about 15 minutes (reproduction is highly conservative—meiosis in all organisms is basically the same). After chromosomal replication, the four available chromatids (the twin copies of the fully replicated chromosomes) are in tight formation with one another. During this time, homologous sites on two chromatids can mesh with one another, and may exchange genetic information. Immediately after replication, the tetrad formed by replication contains two pairs of two identical chromatids; after crossing over, each of the four chromatids carries a unique set of genetic information.
This reordering process, which is not well understood, seems to involve many molecules (RNA and protein recombinases). Enzymes known as recombinases catalyze the reactions that allow for crossover to occur. A recombinase creates a nick in one strand of a DNA double helix, allowing the nicked strand to pull apart from its complementary strand and anneal to one strand of the double helix on the opposite chromatid. A second nick allows the unannealed strand in the second double helix to pull apart and anneal to the remaining strand in the first, forming a structure known as a cross-strand exchange or a "Holliday junction." The Holliday junction is a tetrahedral structure that can be "pulled" by other recombinases, moving it along the four-stranded structure.
Guided by this complex of RNA and proteins, a variable amount of cutting and pasting between these eight strands occurs. The genetic material is organized in a modular fashion, and the shuffling involves these modules. It is currently an open question as to whether this tetraplex reorganization of the genetic inheritance is random as classical science would have it, or is as well organized and well-designed as the ribosome-triplet code method of protein synthesis.
A well-characterized example of such DNA manipulation by RNA/protein complexes is to be found in the make-antibody programming of an immature lymphocyte in the immune system. This is directed variation within a well-defined permutation space. The antibody program carried by the immature lymphocyte is then run in a virtual-body environment generated by the thymus. If the program running in this virtual reality recognizes a body component, the lymphocyte is destroyed; else it is activated and released as a mature T-cell.
This immune system manipulation of DNA modules has been a focus of study as it has great relevance to disease. The DNA manipulation in the tetraplex stage, on the other hand, is not well characterized, but the basic principles are probably similar.
The simplest type of recombination is crossing over, where strands are cut across and then reconnected differently. Even this aspect is highly modular; there are hot spots where crossing over occurs frequently, and forbidden zones where it never happens.
The most complex rearrangement of the genetic modules occurs during speciation. For instance, along the lineage that diverged from the apes to humans, two chromosomes became fused into one, the human Chromosome Two, the second largest one. Such massive changes and manipulation of the genome by the RNA-protein complex are not well understood.
Consequences of crossover
In most eukaryotes, a cell carries two copies of each gene, each DNA coding of the gene referred to as an allele. Each parent passes on one allele to each offspring. Even without recombination, each gamete (sex cell) contains a random assortment of chromatids, choosing randomly from each pair of chromatids available. With recombination, however, the gamete can receive a (mostly) random assortment of individual genes, as each chromosome may contain genetic information from two different chromatids.
Recombination results in a new arrangement of maternal and paternal alleles on the same chromosome. Although the same genes appear in the same order, the alleles are different. This process explains why offspring from the same parents can look so different. In this way, it is theoretically possible to have any combination of parental alleles in an offspring, and the fact that two alleles appear together in one offspring does not have any influence on the statistical probability that another offspring will have the same combination. This theory of "independent assortment" of alleles is fundamental to genetic inheritance. However, there is an exception that requires further discussion.
The frequency of recombination is actually not the same for all gene combinations. This is because recombination is greatly influenced by the proximity of one gene to another. If two genes are located close together on a chromosome, the likelihood that a recombination event will separate these two genes is less than if they were farther apart. Genetic linkage describes the tendency of genes to be inherited together as a result of their location on the same chromosome. Linkage disequilibrium describes a situation in which some combinations of genes or genetic markers occur more or less frequently in a population than would be expected from their distances apart. This concept is applied when searching for a gene that may cause a particular disease. This is done by comparing the occurrence of a specific DNA sequence with the appearance of a disease. When a high correlation between the two is found, it is likely that the appropriate gene sequence is closer.
Other types of DNA reordering (non-sexual)
Crossover recombination can occur between any two double helices of DNA that are very close in sequence and come into contact with one another. Thus, crossover may occur between Alu repeats on the same chromatid, or between similar sequences on two completely different chromosomes. These processes are called unbalanced recombination. Unbalanced recombination is fairly rare compared to normal recombination, but severe problems can arise if a gamete containing unbalanced recombinants becomes part of a zygote. Offspring with severe unbalances rarely live through birth.
In molecular biology, recombination generally refers to the molecular process by which genetic variation found associated at two different places in a continuous piece of DNA becomes disassociated (shuffled). In this process one or both of the genetic variants are replaced by different variants found at same two places in a second DNA molecule. One mechanism leading to such molecular recombination is chromosomal crossing over. Such shuffling of variation is also possible between duplicated loci within the same DNA molecule. If the shuffling process changes the number of loci in each of the recombinant molecules, one speaks of "unbalanced" recombination or unequal crossing over.
In conservative site-specific recombination, a mobile DNA element is inserted into a strand of DNA by means similar to that seen in crossover. A segment of DNA on the mobile element matches exactly with a segment of DNA on the target, allowing enzymes called integrases to insert the rest of the mobile element into the target.
Another form of site-specific recombination, transpositional recombination does not require an identical strand of DNA in the mobile element to match with the target DNA. Instead, the integrases involved introduce nicks in both the mobile element and the target DNA, allowing the mobile DNA to enter the sequence. The nicks are then removed by ligases.
ReferencesISBN links support NWE through referral fees
- Alberts, B. et al., Molecular Biology of the Cell, 3rd Edition. Garland Publishing, 1994.
- Mayerhofer R, Koncz-Kalman Z, Nawrath C, Bakkeren G, Crameri A, Angelis K, Redei GP, Schell J, Hohn B, Koncz C. T-DNA integration: a mode of illegitimate recombination in plants. EMBO J. 1991 Mar;10(3):697-704.
- This article contains material from the Science Primer published by the NCBI, which, as a US government publication, is in the public domain at http://www.ncbi.nlm.nih.gov/About/disclaimer.html.
Credits
New World Encyclopedia writers and editors rewrote and completed the Wikipedia article in accordance with New World Encyclopedia standards. This article abides by terms of the Creative Commons CC-by-sa 3.0 License (CC-by-sa), which may be used and disseminated with proper attribution. Credit is due under the terms of this license that can reference both the New World Encyclopedia contributors and the selfless volunteer contributors of the Wikimedia Foundation. To cite this article click here for a list of acceptable citing formats.The history of earlier contributions by wikipedians is accessible to researchers here:
The history of this article since it was imported to New World Encyclopedia:
Note: Some restrictions may apply to use of individual images which are separately licensed.