Denaturation (biochemistry)
If proteins in a living cell are denatured, this results in disruption of cell activity and possibly cell death. Denatured proteins can exhibit a wide range of characteristics, from loss of solubility to communal aggregation. Denatured alcohol is an exception to this definition, as the term refers not to any alteration of the substance's structure but to the addition of toxins and other things to render it undrinkable.
Common examples
When food is cooked, some of its proteins become denatured. This is why boiled eggs become hard and cooked meat becomes firm.
A classic example of denaturing in proteins comes from egg whites, which are largely egg albumins in water. Fresh from the eggs, egg whites are transparent and liquid. Cooking the thermally unstable whites turns them opaque, forming an interconnected solid mass. The same transformation can be effected with a denaturing chemical. Pouring egg whites into a beaker of acetone will also turn egg whites opaque and solid. The skin which forms on curdled milk is another common example of denatured protein. And the traditional Peruvian cold appetizer known as ceviche is prepared by chemically "cooking" raw fish and shellfish in an acidic citrus marinade, without heat.
Although denaturating egg whites is irreversible, in many other cases, denaturing is reversible.
Denatured proteins can exhibit a wide range of characteristics, from loss of solubility to communal aggregation. Communal aggregation is the phenomenon of aggregation of the hydrophobic proteins to come closer and form the bonding between them, so as to reduce the total area exposed to water. It is a very common problem with the hydrophobic protein to make aggregates. Such aggregates hamper the filtration process by cake formation.
Background
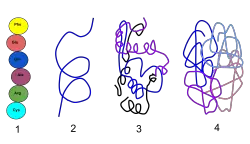
1) Primary Structure : the linear structure of amino acids in the polypeptide chain
2) Secondary Structure : hydrogen bonds between peptide group chains in an alpha helix or beta
3) Tertiary Structure : three-dimensional structure of alpha helixes and beta helixes folded
4) Quaternary Structure : three-dimensional structure of multiple polypeptides and how they fit together
Proteins are very long strands of amino acids linked together in specific sequences. A protein is created by ribosomes that "read" mRNA that is encoded by codons in the gene and assemble the requisite amino acid combination from the genetic instruction, in a process known as translation. The newly created protein strand then undergoes posttranslational modification, in which additional atoms or molecules are added, for example copper, zinc or iron. Once this post-translational modification process has been completed, the protein begins to fold (spontaneously, and sometimes with enzymatic assistance), curling up on itself so that hydrophobic elements of the protein are buried deep inside the structure and hydrophilic elements end up on the outside. The final shape of a protein determines how it interacts with its environment.
When a protein is denatured, the secondary and tertiary structures are altered but the peptide bonds between the amino acids are left intact. Since the structure of the protein determines its function, the protein can no longer perform its function once it has been denatured. This is in contrast to intrinsically unstructured proteins, which are unfolded in their native state, but still functionally active.
How denaturation occurs at levels of protein structure
- In quaternary structure denaturation, protein sub-units are dissociated and/or the spatial arrangement of protein subunits is disrupted.
- Tertiary structure denaturation involves the disruption of:
- In secondary structure denaturation, proteins lose all regular repeating patterns such as alpha-helices and beta-pleated sheets, and adopt a random coil configuration.
- Primary structure, such as the sequence of amino acids held together by covalent peptide bonds, is not disrupted by denaturation.
Loss of function
Most biological proteins lose their biological function when denatured. For example, enzymes lose their activity, because the substrates can no longer bind to the active site, and because amino acid residues involved in stabilizing substrates' transition states are no longer positioned to be able to do so.
Reversibility and irreversibility
In many proteins (unlike egg whites), denaturation is reversible (the proteins can regain their native state when the denaturing influence is removed). This was important historically, as it led to the notion that all the information needed for proteins to assume their native state was encoded in the primary structure of the protein, and hence in the DNA that codes for the protein.
Nucleic acid denaturation
The denaturation of nucleic acids such as DNA due to high temperatures, is the separation of a double strand into two single strands, which occurs when the hydrogen bonds between the strands are broken. This may occur during polymerase chain reaction. Nucleic acid strands realign when "normal" conditions are restored during annealing. If the conditions are restored too quickly, the nucleic acid strands may realign imperfectly.
Denaturants
Acids
Acidic protein denaturants include:
- Acetic acid
- Trichloroacetic acid 12% in water
- Sulfosalicylic acid
Solvents
Most organic solvents are denaturing, including:
Cross linking reagents
Cross linking agents for proteins include:
- Formaldehyde
- Glutaraldehyde
Chaotropic agents
Chaotropic agents include:
- Urea 6 - 8 mol/l
- Guanidinium chloride 6 mol/l
- Lithium perchlorate 4.5 mol/l
Disulfide bond reducers
Agents that break disulfide bonds by reduction include:
- 2-Mercaptoethanol
- Dithiothreitol
- TCEP (tris(2-carboxyethyl)phosphine)
Other
- Picric acid
See also
ReferencesISBN links support NWE through referral fees
- Lapanje, Savo. 1978. Physicochemical Aspects of Protein Denaturation. Wiley. ISBN 978-0471034094
- MacNaught, Alan D., and Andrew R. Wilkinson (eds.). 1997. Compendium of Chemical Terminology: IUPAC Recommendations (the "Gold Book"). Blackwell Science. ISBN 0865426848
- Whitford, David. 2005. Proteins: Structure and Function. Wiley. ISBN 978-0471498940
Credits
New World Encyclopedia writers and editors rewrote and completed the Wikipedia article in accordance with New World Encyclopedia standards. This article abides by terms of the Creative Commons CC-by-sa 3.0 License (CC-by-sa), which may be used and disseminated with proper attribution. Credit is due under the terms of this license that can reference both the New World Encyclopedia contributors and the selfless volunteer contributors of the Wikimedia Foundation. To cite this article click here for a list of acceptable citing formats.The history of earlier contributions by wikipedians is accessible to researchers here:
The history of this article since it was imported to New World Encyclopedia:
Note: Some restrictions may apply to use of individual images which are separately licensed.

